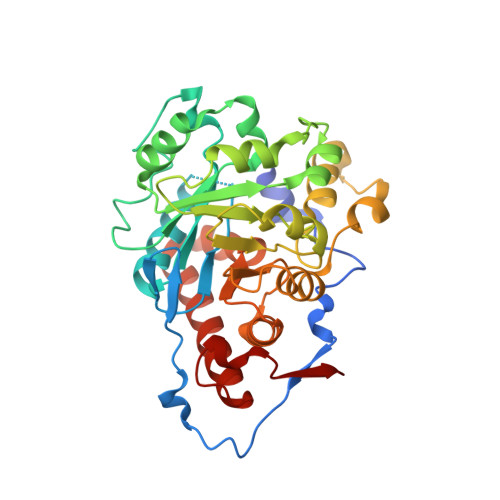
Crystal structure of type 2 isopentenyl diphosphate isomerase from Thermus thermophilus in complex with inorganic pyrophosphate
de Ruyck, J., Pouyez, J., Rothman, S.C., Poulter, D., Wouters, J.(2008) Biochemistry 47: 9051-9053
- PubMed: 18693754 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi801159x
- Primary Citation Related Structures:
3DH7 - PubMed Abstract:
The N-terminal region is stabilized in the crystal structure of Thermus thermophilus type 2 isopentenyl diphosphate isomerase in complex with inorganic pyrophosphate, providing new insights about the active site and the catalytic mechanism of the enzyme. The PP i moiety is located near the conserved residues, H10, R97, H152, Q157, E158, and W219, and the flavin cofactor. The putative active site of isopentenyl diphosphate isomerase 2 provides interactions for stabilizing a carbocationic intermediate similar to those that stabilize the intermediate in the well-established protonation-deprotonation mechanism of isopentenyl diphosphate isomerase 1.
- Department of Chemistry, University of Namur, Namur, Belgium. jerome.deruyck@fundp.ac.be
Organizational Affiliation: