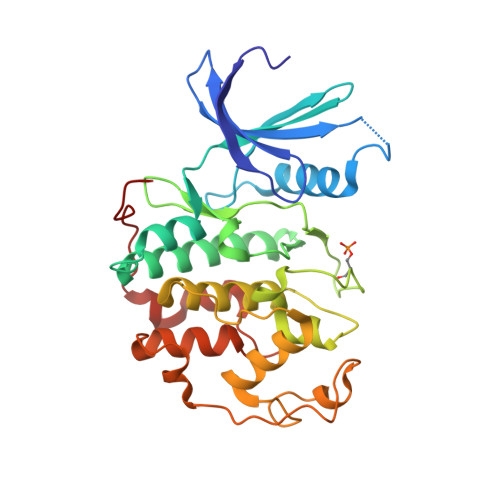
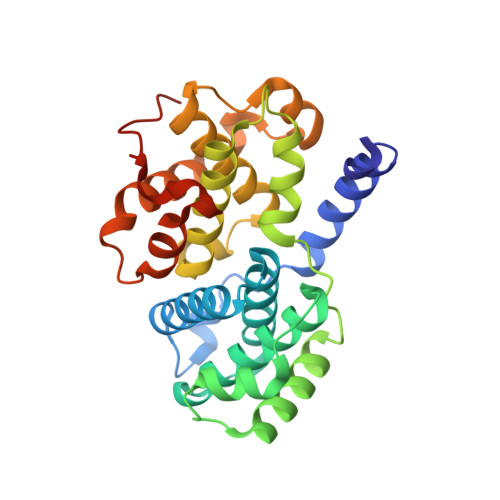
CR8, a potent and selective, roscovitine-derived inhibitor of cyclin-dependent kinases.
Bettayeb, K., Oumata, N., Echalier, A., Ferandin, Y., Endicott, J.A., Galons, H., Meijer, L.(2008) Oncogene 27: 5797-5807
- PubMed: 18574471 Search on PubMed
- DOI: https://doi.org/10.1038/onc.2008.191
- Primary Citation Related Structures:
3DDP, 3DDQ - PubMed Abstract:
Among the ten pharmacological inhibitors of cyclin-dependent kinases (CDKs) currently in clinical trials, the purine roscovitine (CYC202, Seliciclib) is undergoing phase 2 trials against non-small-cell lung and nasopharyngeal cancers. An extensive medicinal chemistry study, designed to generate more potent analogues of roscovitine, led to the identification of an optimal substitution at the N6 position (compound CR8). An extensive selectivity study (108 kinases) highlights the exquisite selectivity of CR8 for CDK1/2/3/5/7/9. CR8 was 2- to 4-fold more potent than (R)-roscovitine at inhibiting these kinases. Cocrystal structures of (R)-CR8 and (R)-roscovitine with pCDK2/cyclin A showed that both inhibitors adopt essentially identical positions. The cellular effects of CR8 and (R)-roscovitine were investigated in human neuroblastoma SH-SY5Y cells. CR8 inhibited the phosphorylation of CDK1 and 9 substrates, with a 25-50 times higher potency compared to (R)-roscovitine. CR8 was consistently more potent than (R)-roscovitine at inducing apoptotic cell death parameters: 3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium reduction (40-fold), lactate dehydrogenase release (35-fold), caspases activation (68-fold) and poly-(ADP-ribose)polymerase cleavage (50-fold). This improved cell death-inducing activity of CR8 over (R)-roscovitine was observed in 25 different cell lines. Altogether these results show that second-generation analogues of (R)-roscovitine can be designed with improved antitumor potential.
- CNRS, Cell Cycle Group, Station Biologique, Bretagne, France.
Organizational Affiliation: