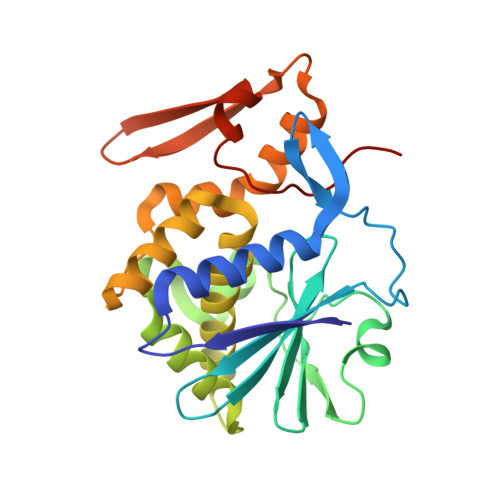
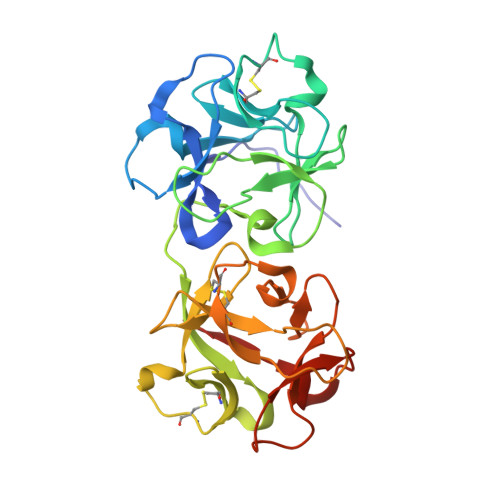
Structure of mistletoe lectin I from Viscum album in complex with the phytohormone zeatin
Meyer, A., Rypniewski, W., Szymanski, M., Voelter, W., Barciszewski, J., Betzel, C.(2008) Biochim Biophys Acta 1784: 1590-1595
- PubMed: 18718563 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2008.07.010
- Primary Citation Related Structures:
3D7W - PubMed Abstract:
The crystal structure of mistletoe lectin I (ML-I) isolated from the European mistletoe Viscum album in complex with the most active phytohormone zeatin has been analyzed and refined to 2.54 A resolution. X-ray suitable crystals of ML-I were obtained by the counter-diffusion method using the Gel-Tube R crystallization kit (GT-R) onboard the Russian Service Module on the international space station ISS. High quality hexagonal bipyramidal crystals were grown during 3 months under microgravity conditions. Selected crystals were soaked in a saturated solution of zeatin and subsequently diffraction data were collected applying synchrotron radiation. A distinct F(o)-F(c) electron density has been found inside a binding pocket located in subunit B of ML-I and has been interpreted as a single zeatin molecule. The structure was refined to investigate the zeatin-ML-I interactions in detail. The results demonstrate the ability of mistletoe to protect itself from the host transpiration regulation by absorbing the most active host plant hormones as part of a defense mechanism.
- Institute of Biochemistry and Molecular Biology, University of Hamburg, c/o DESY, Notkestr. 85, Building 22a, 22603 Hamburg, Germany.
Organizational Affiliation: