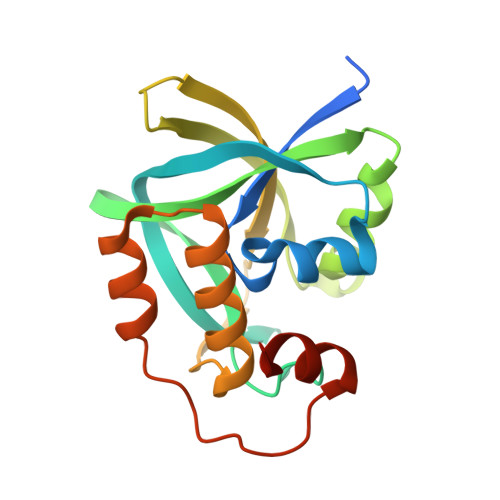
The identification and structural characterization of C7orf24 as gamma-glutamyl cyclotransferase. An essential enzyme in the gamma-glutamyl cycle
Oakley, A.J., Yamada, T., Liu, D., Coggan, M., Clark, A.G., Board, P.G.(2008) J Biological Chem 283: 22031-22042
- PubMed: 18515354 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M803623200
- Primary Citation Related Structures:
2PN7, 2RBH, 3CRY - PubMed Abstract:
The hypothetical protein C7orf24 has been implicated as a cancer marker with a potential role in cell proliferation. We have identified C7orf24 as gamma-glutamyl cyclotransferase (GGCT) that catalyzes the formation of 5-oxoproline (pyroglutamic acid) from gamma-glutamyl dipeptides and potentially plays a significant role in glutathione homeostasis. In the present study we have identified the first cDNA clones encoding a gamma-glutamyl cyclotransferase. The GGCT gene is located on chromosome 7p14-15 and consists of four exons that span 8 kb. The primary sequence is 188 amino acids in length and is unlike any protein of known function. We crystallized functional recombinant gamma-glutamyl cyclotransferase and determined its structure at 1.7 A resolution. The enzyme is a dimer of 20,994-Da subunits. The topology of GGCT is unrelated to other enzymes associated with cyclotransferase-like activity. The fold was originally classified as "BtrG-like," a small family that only includes structures of hypothetical proteins from Mus musculus, Escherichia coli, Pyrococcus horikoshii, and Arabidopsis thaliana. Since this is the first member of this family with a defined function, we propose to refer to this structure as the gamma-glutamyl cyclotransferase fold. We have identified a potential active site pocket that contains a highly conserved glutamic acid (Glu(98)) and propose that it acts as a general acid/base in the reaction mechanism. Mutation of Glu(98) to Ala or Gln completely inactivates the enzyme without altering the overall fold.
- Research School of Chemistry, Australian National University, Canberra, Australian Capital Territory 2601, Australia.
Organizational Affiliation: