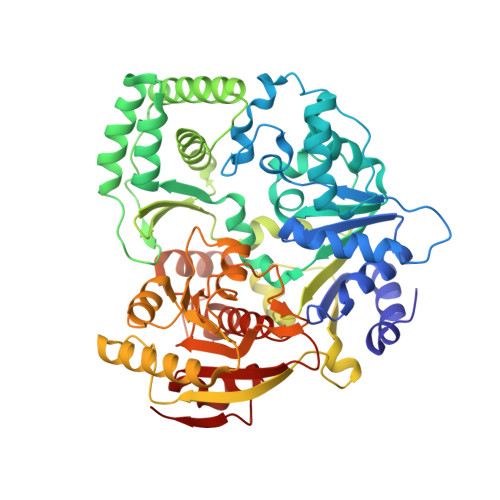
XPD helicase structures and activities: insights into the cancer and aging phenotypes from XPD mutations.
Fan, L., Fuss, J.O., Cheng, Q.J., Arvai, A.S., Hammel, M., Roberts, V.A., Cooper, P.K., Tainer, J.A.(2008) Cell 133: 789-800
- PubMed: 18510924 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2008.04.030
- Primary Citation Related Structures:
3CRV, 3CRW - PubMed Abstract:
Mutations in XPD helicase, required for nucleotide excision repair (NER) as part of the transcription/repair complex TFIIH, cause three distinct phenotypes: cancer-prone xeroderma pigmentosum (XP), or aging disorders Cockayne syndrome (CS), and trichothiodystrophy (TTD). To clarify molecular differences underlying these diseases, we determined crystal structures of the XPD catalytic core from Sulfolobus acidocaldarius and measured mutant enzyme activities. Substrate-binding grooves separate adjacent Rad51/RecA-like helicase domains (HD1, HD2) and an arch formed by 4FeS and Arch domains. XP mutations map along the HD1 ATP-binding edge and HD2 DNA-binding channel and impair helicase activity essential for NER. XP/CS mutations both impair helicase activity and likely affect HD2 functional movement. TTD mutants lose or retain helicase activity but map to sites in all four domains expected to cause framework defects impacting TFIIH integrity. These results provide a foundation for understanding disease consequences of mutations in XPD and related 4Fe-4S helicases including FancJ.
- Department of Molecular Biology, Skaggs Institute of Chemical Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: