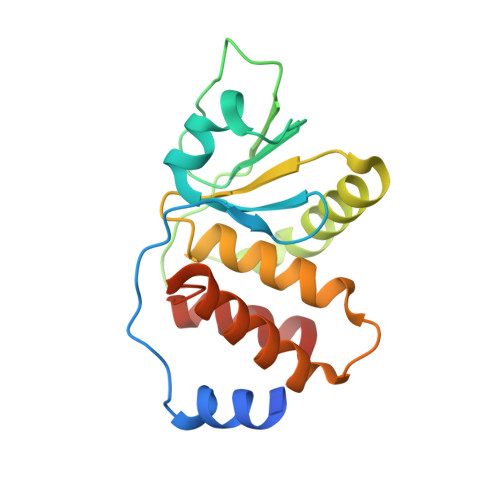
Dimeric Quaternary Structure of the Prototypical Dual Specificity Phosphatase VH1.
Koksal, A.C., Nardozzi, J.D., Cingolani, G.(2009) J Biological Chem 284: 10129-10137
- PubMed: 19211553 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M808362200
- Primary Citation Related Structures:
3CM3 - PubMed Abstract:
The Vaccinia virus H1 gene product, VH1, is a dual specificity phosphatase that down-regulates the cellular antiviral response by dephosphorylating STAT1. The crystal structure of VH1, determined at 1.32 A resolution, reveals a novel dimeric quaternary structure, which exposes two active sites spaced approximately 39 A away from each other. VH1 forms a stable dimer via an extensive domain swap of the N-terminal helix (residues 1-20). In vitro, VH1 can dephosphorylate activated STAT1, in a reaction that is competed by the nuclear transport adapter importin alpha5. Interestingly, VH1 is inactive with respect to STAT1 bound to DNA, suggesting that the viral phosphatase acts predominantly on the cytoplasmic pool of activated STAT1. We propose that the dimeric quaternary structure of VH1 is essential for specific recognition of activated STAT1, which prevents its nuclear translocation, thus blocking interferon-gamma signal transduction and antiviral response.
- Department of Biochemistry and Molecular Biology, SUNY Upstate Medical University, Syracuse, New York 13210, USA.
Organizational Affiliation: