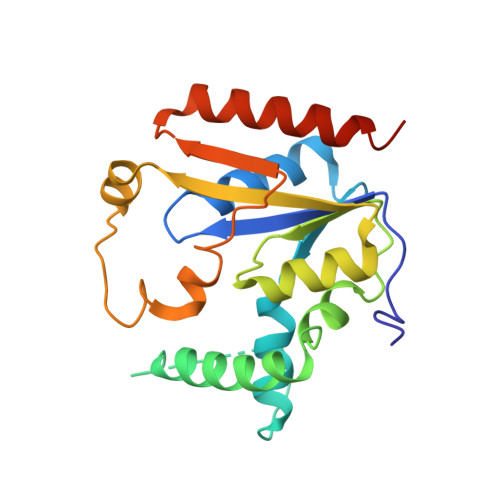
Crystal structure of human phosphomavelonate kinase at 1.8 A resolution
Chang, Q., Yan, X.-X., Gu, S.-Y., Liu, J.-F., Liang, D.-C.(2008) Proteins 73: 254-258
- PubMed: 18618710 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22151
- Primary Citation Related Structures:
3CH4 - National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, People's Republic of China.
Organizational Affiliation: