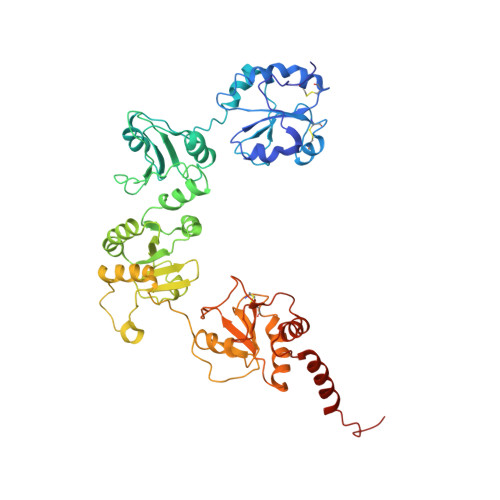
The Catalytic Activity of Protein-disulfide Isomerase Requires a Conformationally Flexible Molecule.
Tian, G., Kober, F.X., Lewandrowski, U., Sickmann, A., Lennarz, W.J., Schindelin, H.(2008) J Biological Chem 283: 33630-33640
- PubMed: 18815132 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M806026200
- Primary Citation Related Structures:
3BOA - PubMed Abstract:
Protein-disulfide isomerase (PDI) catalyzes the formation of the correct pattern of disulfide bonds in secretory proteins. A low resolution crystal structure of yeast PDI described here reveals large scale conformational changes compared with the initially reported structure, indicating that PDI is a highly flexible molecule with its catalytic domains, a and a', representing two mobile arms connected to a more rigid core composed of the b and b' domains. Limited proteolysis revealed that the linker between the a domain and the core is more susceptible to degradation than that connecting the a' domain to the core. By restricting the two arms with inter-domain disulfide bonds, the molecular flexibility of PDI, especially that of its a domain, was demonstrated to be essential for the enzymatic activity in vitro and in vivo. The crystal structure also featured a PDI dimer, and a propensity to dimerize in solution and in the ER was confirmed by cross-linking experiments and the split green fluorescent protein system. Although sedimentation studies suggested that the self-association of PDI is weak, we hypothesize that PDI exists as an interconvertible mixture of monomers and dimers in the endoplasmic reticulum due to its high abundance in this compartment.
- Department of Biochemistry and Cell Biology, Stony Brook University, Stony Brook, New York 11794-5215, USA.
Organizational Affiliation: