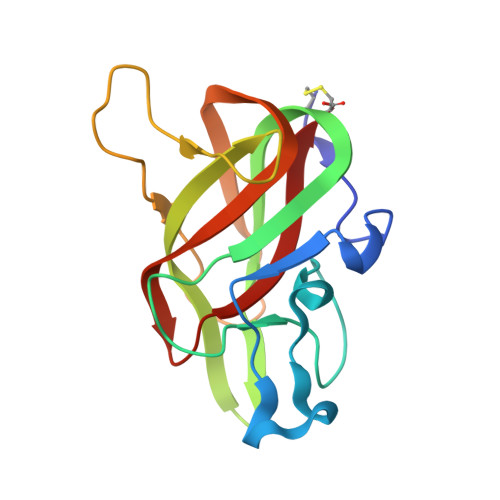
Crystal structure of lactadherin C2 domain at 1.7A resolution with mutational and computational analyses of its membrane-binding motif.
Shao, C., Novakovic, V.A., Head, J.F., Seaton, B.A., Gilbert, G.E.(2008) J Biological Chem 283: 7230-7241
- PubMed: 18160406 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M705195200
- Primary Citation Related Structures:
3BN6 - PubMed Abstract:
Lactadherin is a phosphatidyl-L-serine (Ptd-L-Ser)-binding protein that decorates membranes of milk fat globules. The major Ptd-l-Ser binding function of lactadherin has been localized to its C2 domain, which shares homology with the C2 domains of blood coagulation factor VIII and factor V. Correlating with this homology, purified lactadherin competes efficiently with factors VIII and V for Ptd-L-Ser binding sites, functioning as a potent anticoagulant. We have determined the crystal structure of the lactadherin C2 domain (Lact-C2) at 1.7A resolution. The bovine Lact-C2 structure has a beta-barrel core that is homologous with the factor VIII C2 (fVIII-C2) and factor V C2 (fV-C2) domains. Two loops at the end of the beta-barrel, designated spikes 1 and 3, display four water-exposed hydrophobic amino acids, reminiscent of the membrane-interactive residues of fVIII-C2 and fV-C2. In contrast to the corresponding loops in fVIII-C2 and fV-C2, spike 1 of Lact-C2 adopts a hairpin turn in which the 7-residue loop is stabilized by internal hydrogen bonds. Further, central glycine residues in two membrane-interactive loops may enhance conformability of Lact-C2 to membrane binding sites. Mutagenesis studies confirmed a membrane-interactive role for the hydrophobic and/or Gly residues of both spike 1 and spike 3. Substitution of spike 1 of fVIII-C2 into Lact-C2 also diminished binding. Computational ligand docking studies identified two prospective Ptd-l-Ser interaction sites. These results identify two membrane-interactive loops of Lact-C2 and provide a structural basis for the more efficient phospholipid binding of lactadherin as compared with factor VIII and factor V.
- Department of Physiology and Biophysics, Boston University School of Medicine, Boston, Massachusetts 02118, USA.
Organizational Affiliation: