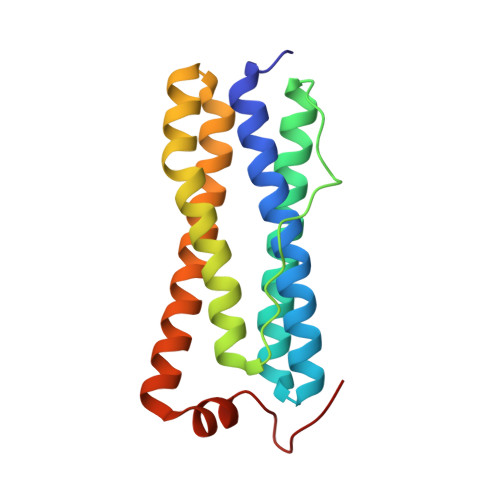
Bacterioferritin from Mycobacterium smegmatis contains zinc in its di-nuclear site.
Janowski, R., Auerbach-Nevo, T., Weiss, M.S.(2008) Protein Sci 17: 1138-1150
- PubMed: 18445621 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.034819.108
- Primary Citation Related Structures:
3BKN - PubMed Abstract:
Bacterioferritins, also known as cytochrome b (1), are oligomeric iron-storage proteins consisting of 24 identical amino acid chains, which form spherical particles consisting of 24 subunits and exhibiting 432 point-group symmetry. They contain one haem b molecule at the interface between two subunits and a di-nuclear metal binding center. The X-ray structure of bacterioferritin from Mycobacterium smegmatis (Ms-Bfr) was determined to a resolution of 2.7 A in the monoclinic space group C2. The asymmetric unit of the crystals contains 12 protein molecules: five dimers and two half-dimers located along the crystallographic twofold axis. Unexpectedly, the di-nuclear metal binding center contains zinc ions instead of the typically observed iron ions in other bacterioferritins.
- EMBL Hamburg Outstation, D-22603 Hamburg, Germany.
Organizational Affiliation: