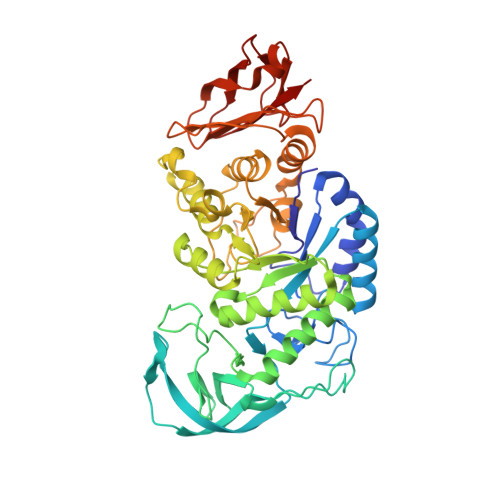
Structure of Bacillus amyloliquefaciens alpha-amylase at high resolution: implications for thermal stability.
Alikhajeh, J., Khajeh, K., Ranjbar, B., Naderi-Manesh, H., Lin, Y.H., Liu, E., Guan, H.H., Hsieh, Y.C., Chuankhayan, P., Huang, Y.C., Jeyaraman, J., Liu, M.Y., Chen, C.J.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 121-129
- PubMed: 20124706 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309109051938
- Primary Citation Related Structures:
3BH4 - PubMed Abstract:
The crystal structure of Bacillus amyloliquefaciens alpha-amylase (BAA) at 1.4 A resolution revealed ambiguities in the thermal adaptation of homologous proteins in this family. The final model of BAA is composed of two molecules in a back-to-back orientation, which is likely to be a consequence of crystal packing. Despite a high degree of identity, comparison of the structure of BAA with those of other liquefying-type alpha-amylases indicated moderate discrepancies at the secondary-structural level. Moreover, a domain-displacement survey using anisotropic B-factor and domain-motion analyses implied a significant contribution of domain B to the total flexibility of BAA, while visual inspection of the structure superimposed with that of B. licheniformis alpha-amylase (BLA) indicated higher flexibility of the latter in the central domain A. Therefore, it is suggested that domain B may play an important role in liquefying alpha-amylases, as its rigidity offers a substantial improvement in thermostability in BLA compared with BAA.
- Life Science Group, Scientific Research Division, National Synchrotron Radiation Research Center, Hsinchu 30076, Taiwan. khajeh@modares.ac.ir
Organizational Affiliation: