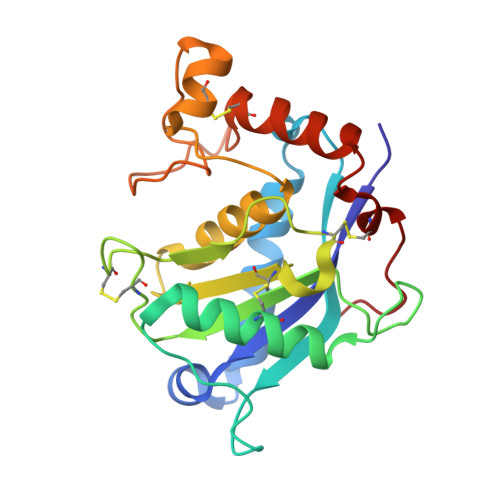
High resolution crystal structure of the catalytic domain of ADAMTS-5 (aggrecanase-2).
Shieh, H.S., Mathis, K.J., Williams, J.M., Hills, R.L., Wiese, J.F., Benson, T.E., Kiefer, J.R., Marino, M.H., Carroll, J.N., Leone, J.W., Malfait, A.M., Arner, E.C., Tortorella, M.D., Tomasselli, A.(2008) J Biological Chem 283: 1501-1507
- PubMed: 17991750 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M705879200
- Primary Citation Related Structures:
3B8Z - PubMed Abstract:
Aggrecanase-2 (a disintegrin and metalloproteinase with thrombospondin motifs-5 (ADAMTS-5)), a member of the ADAMTS protein family, is critically involved in arthritic diseases because of its direct role in cleaving the cartilage component aggrecan. The catalytic domain of aggrecanase-2 has been refolded, purified, and crystallized, and its three-dimensional structure determined to 1.4A resolution in the presence of an inhibitor. A high resolution structure of an ADAMTS/aggrecanase protein provides an opportunity for the development of therapeutics to treat osteoarthritis.
- Pfizer Global Research and Development, St. Louis, Missouri 63017.
Organizational Affiliation: