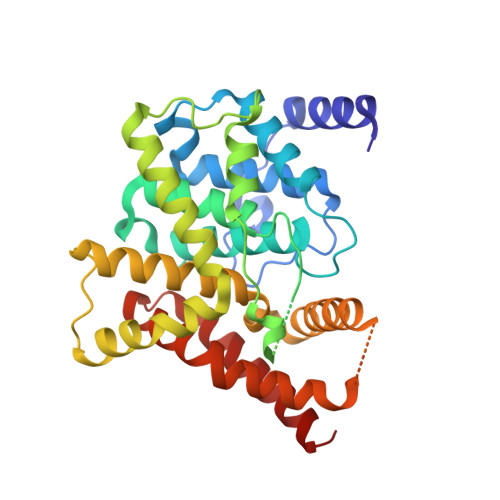
Conformational variations of both phosphodiesterase-5 and inhibitors provide the structural basis for the physiological effects of vardenafil and sildenafil.
Wang, H., Ye, M., Robinson, H., Francis, S.H., Ke, H.(2008) Mol Pharmacol 73: 104-110
- PubMed: 17959709 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1124/mol.107.040212
- Primary Citation Related Structures:
3B2R - PubMed Abstract:
Vardenafil has higher affinity to phosphodiesterase-5 (PDE5) than sildenafil and lower administered dosage for the treatment of erectile dysfunction. However, the molecular basis for these differences is puzzling because two drugs have similar chemical structures. Reported here is a crystal structure of the fully active and nonmutated PDE5A1 catalytic domain in complex with vardenafil. The structure shows that the conformation of the H-loop in the PDE5A1-vardenafil complex is different from those of any known structures of the unliganded PDE5 and its complexes with the inhibitors. In addition, the molecular configuration of vardenafil differs from that of sildenafil when bound to PDE5. It is noteworthy that the binding of vardenafil causes loss of the divalent metal ions that have been observed in all the previously published PDE structures. The conformational variation of both PDE5 and the inhibitors provides structural insight into the different potencies of the drugs.
- Department of Biochemistry and Biophysics, The University of North Carolina, Chapel Hill, NC 27599-7260, USA.
Organizational Affiliation: