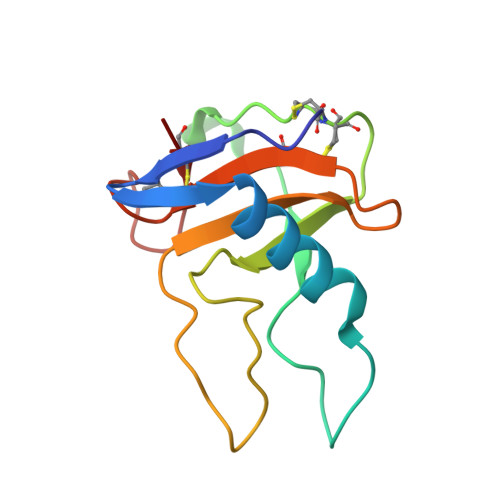
Conformational variation revealed by the crystal structure of RNase U2A complexed with Ca ion and 2'-adenylic acid at 1.03 angstrom resolution.
Noguchi, S.(2010) Protein Pept Lett 17: 1559-1561
- PubMed: 20858208 Search on PubMed
- DOI: https://doi.org/10.2174/0929866511009011559
- PubMed Abstract:
Asparagine can be non-enzymatically deamidated and isomerized via succinimide to isoaspartate. This post-translational modification can potentially alter the physical properties or the function of the parent protein. Asn32 of ribonuclease U2A from Ustilago sphaerogena is known to rapidly deamidate and isomerize in alkaline conditions. The crystal structure of ribonuclease U2A complexed with 2'-adenylic acid and calcium ions was determined at 1.03 Å resolution. In this structure, the region from Asp29 to Asp37 winds around a calcium ion, and the main-chain of Asn32-Gly33 adopts an extended conformation. Rotation of the side-chain of Asn32 could bring Asn32C(γ) into close proximity to Gly33N, in a conformation suitable for succinimide formation. The structure suggests that in solution the region around Asn32-Gly33 is likely to be in equilibrium between multiple conformers, with the deamidation of Asn32 proceeding when the region adopts an extended conformation.
- Graduate School of Pharmaceutical Sciences, The University of Tokyo, Tokyo 113-0033, Japan. snoguchi@mol.f.u-tokyo.ac.jp
Organizational Affiliation: