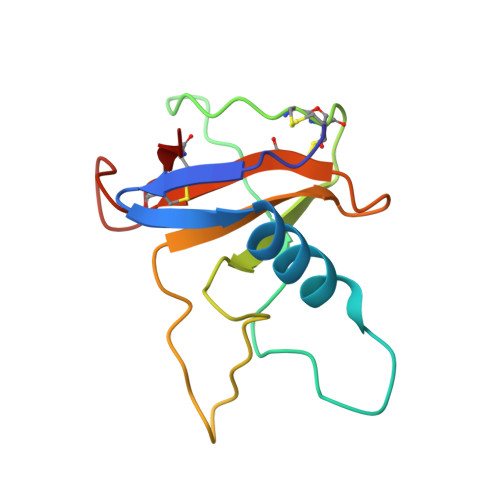
Structural changes induced by the deamidation and isomerization of asparagine revealed by the crystal structure of Ustilago sphaerogena ribonuclease U2B
Noguchi, S.(2010) Biopolymers 93: 1003-1010
- PubMed: 20623666 Search on PubMed
- DOI: https://doi.org/10.1002/bip.21514
- Primary Citation Related Structures:
3AHS - PubMed Abstract:
Under physiological conditions, the deamidation and isomerization of asparagine to isoaspartate (isoAsp) proceeds nonenzymatically via succinimide. Although a large number of proteins have been reported to contain isoAsp, information concerning the three-dimensional structure of proteins containing isoaspartate is still limited. We have crystallized isoAsp containing Ustilago sphaerogena ribonuclease U2B, and determined the crystal structure at 1.32 Å resolution. The structure revealed that the formation of isoAsp32 induces a single turn unfolding of the α-helix from Asp29 to Asp34, and the region from Asp29 to Arg35 forms a U-shaped loop structure. The electron density map shows that isoAsp32 retained the L-configuration at the C(α) atom. IsoAsp32 is in gauche conformation about a C(α)--C(β) bond, and the polypeptide chain bends by ∼90° at isoAsp32. IsoAsp32 protrudes from the surface of the protein, and the abnormal β-peptide bond in the main-chain and α-carboxylate in the side-chain is fully exposed. The structure suggests that the deamidation of the Asn and the isoAsp formation in proteins could confer immunogenicity.
- Graduate School of Pharmaceutical Sciences, The University of Tokyo, Bunkyo-Ku, Tokyo 113-0033, Japan. snoguchi@mol.f.u-tokyo.ac.jp
Organizational Affiliation: