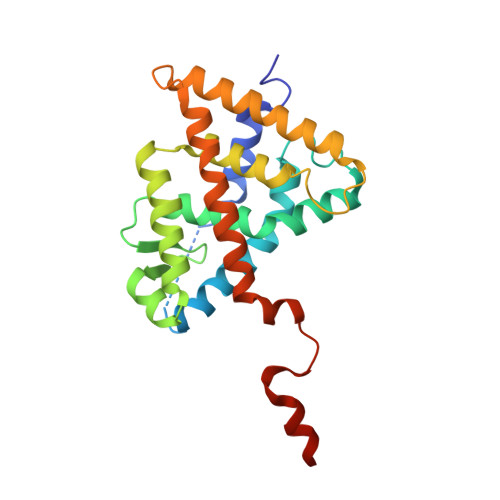
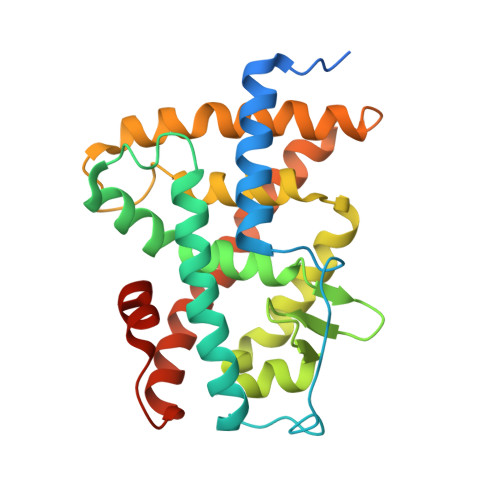
The Phantom Effect of the Rexinoid LG100754: structural and functional insights
Sato, Y., Ramalanjaona, N., Huet, T., Potier, N., Osz, J., Antony, P., Peluso-Iltis, C., Poussin-Courmontagne, P., Ennifar, E., Mely, Y., Dejaegere, A., Moras, D., Rochel, N.(2010) PLoS One 5: e15119-e15119
- PubMed: 21152046 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0015119
- Primary Citation Related Structures:
3A9E - PubMed Abstract:
Retinoic acid receptors (RARs) and Retinoid X nuclear receptors (RXRs) are ligand-dependent transcriptional modulators that execute their biological action through the generation of functional heterodimers. RXR acts as an obligate dimer partner in many signalling pathways, gene regulation by rexinoids depending on the liganded state of the specific heterodimeric partner. To address the question of the effect of rexinoid antagonists on RAR/RXR function, we solved the crystal structure of the heterodimer formed by the ligand binding domain (LBD) of the RARα bound to its natural agonist ligand (all-trans retinoic acid, atRA) and RXRα bound to a rexinoid antagonist (LG100754). We observed that RARα exhibits the canonical agonist conformation and RXRα an antagonist one with the C-terminal H12 flipping out to the solvent. Examination of the protein-LG100754 interactions reveals that its propoxy group sterically prevents the H12 associating with the LBD, without affecting the dimerization or the active conformation of RAR. Although LG100754 has been reported to act as a 'phantom ligand' activating RAR in a cellular context, our structural data and biochemical assays demonstrate that LG100754 mediates its effect as a full RXR antagonist. Finally we show that the 'phantom ligand effect' of the LG100754 is due to a direct binding of the ligand to RAR that stabilizes coactivator interactions thus accounting for the observed transcriptional activation of RAR/RXR.
- Département de Biologie et de Génomique Structurales, Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC), Institut National de Santé et de Recherche Médicale (INSERM) U964/Centre National de Recherche Scientifique (CNRS) UMR 1704/Université de Strasbourg, Illkirch, France.
Organizational Affiliation: