Structural determinant for switching between the polymerase and exonuclease modes in the PCNA-replicative DNA polymerase complex
Nishida, H., Mayanagi, K., Kiyonari, S., Sato, Y., Ishino, Y., Morikawa, K.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA polymerase | 775 | Pyrococcus furiosus | Mutation(s): 0 Gene Names: pol, PF0212 EC: 2.7.7.7 | 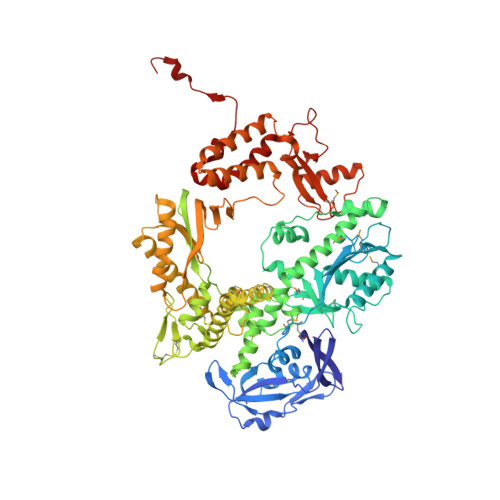 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P61875 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA polymerase sliding clamp | 248 | Pyrococcus furiosus | Mutation(s): 3 Gene Names: pcn, PF0983 | 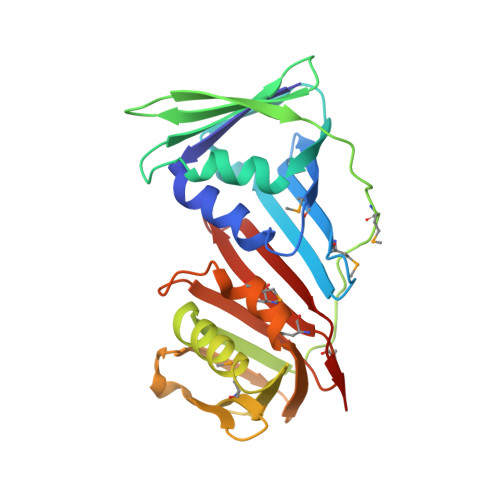 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O73947 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MSE Query on MSE | A | L-PEPTIDE LINKING | C5 H11 N O2 Se |  | MET |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 77.354 | α = 90 |
| b = 90.451 | β = 90 |
| c = 186.211 | γ = 90 |
| Software Name | Purpose |
|---|---|
| CNS | refinement |
| HKL-2000 | data collection |
| HKL-2000 | data reduction |
| SCALEPACK | data scaling |
| SOLVE | phasing |