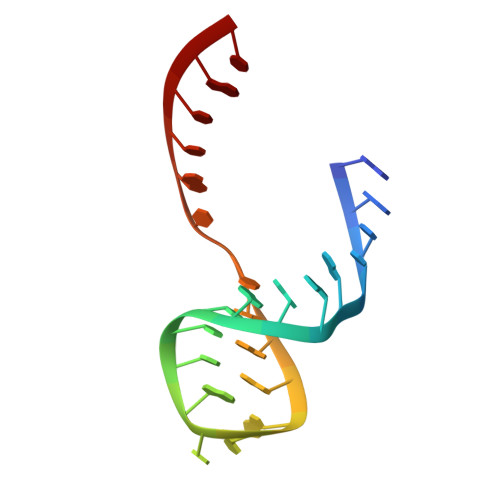
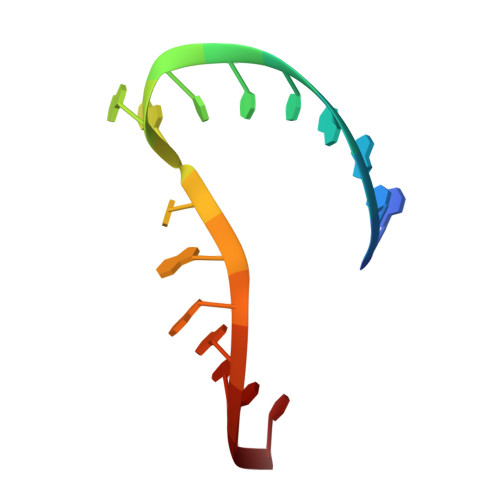Inhibition of the hammerhead ribozyme cleavage reaction by site-specific binding of Tb.
Feig, A.L., Scott, W.G., Uhlenbeck, O.C.(1998) Science 279: 81-84
- PubMed: 9417029 Search on PubMed
- DOI: https://doi.org/10.1126/science.279.5347.81
- Primary Citation Related Structures:
359D - PubMed Abstract:
Terbium(III) [Tb(III)] was shown to inhibit the hammerhead ribozyme by competing with a single magnesium(II) ion. X-ray crystallography revealed that the Tb(III) ion binds to a site adjacent to an essential guanosine in the catalytic core of the ribozyme, approximately 10 angstroms from the cleavage site. Synthetic modifications near this binding site yielded an RNA substrate that was resistant to Tb(III) binding and capable of being cleaved, even in the presence of up to 20 micromolar Tb(III). It is suggested that the magnesium(II) ion thought to bind at this site may act as a switch, affecting the conformational changes required to achieve the transition state.
- Department of Chemistry and Biochemistry, University of Colorado, Boulder, CO 80309, USA.
Organizational Affiliation: