Crystal structure of cytochrome c554 from Vibrio parahaemolyticus strain RIMD2210633
Akazaki, H., Kawai, F., Kumaki, Y., Sekine, K., Hakamata, W., Nishio, T., Park, S.-Y., Oku, T.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cytochrome c554 | 103 | Vibrio parahaemolyticus RIMD 2210633 | Mutation(s): 0 Gene Names: VP2300 | 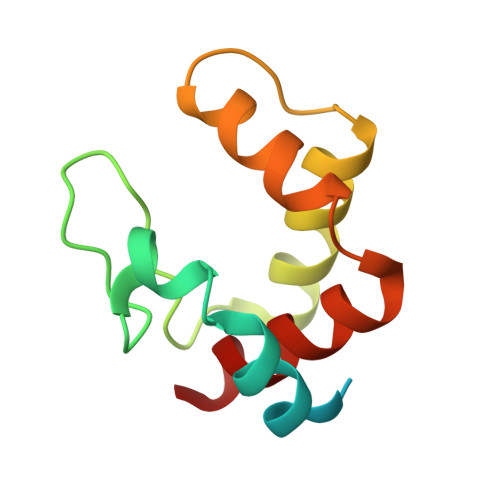 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q87MF5 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| HEC Download:Ideal Coordinates CCD File | AB [auth R] CB [auth S] DB [auth T] EB [auth U] FB [auth V] | HEME C C34 H34 Fe N4 O4 HXQIYSLZKNYNMH-LJNAALQVSA-N |  | ||
| GOL Download:Ideal Coordinates CCD File | BB [auth R], JA [auth C], MA [auth E], YA [auth P] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 84.949 | α = 71.5 |
| b = 87.612 | β = 72.98 |
| c = 103.847 | γ = 83.68 |
| Software Name | Purpose |
|---|---|
| ADSC | data collection |
| MOLREP | phasing |
| REFMAC | refinement |
| HKL-2000 | data reduction |
| SCALEPACK | data scaling |