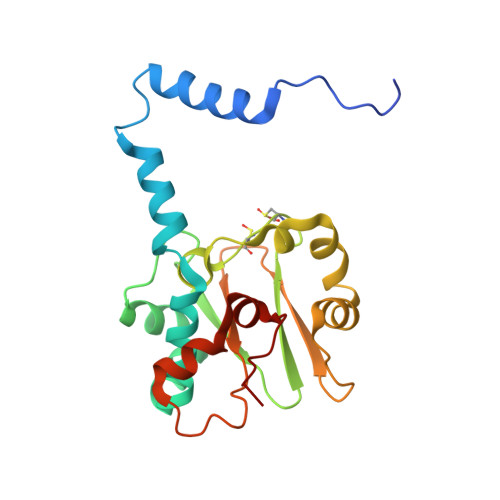
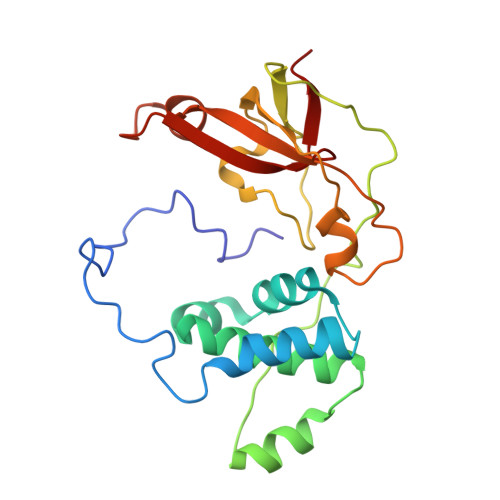
Mutational study on alphaGln90 of Fe-type nitrile hydratase from Rhodococcus sp. N771
Takarada, H., Kawano, Y., Hashimoto, K., Nakayama, H., Ueda, S., Yohda, M., Kamiya, N., Dohmae, N., Maeda, M., Odaka, M.(2006) Biosci Biotechnol Biochem 70: 881-889
- PubMed: 16636455 Search on PubMed
- DOI: https://doi.org/10.1271/bbb.70.881
- Primary Citation Related Structures:
2ZCF - PubMed Abstract:
Nitrile hydratase (NHase) from Rhodococcus sp. N771 is a non-heme iron enzyme having post-translationally modified cysteine ligands, alphaCys112-SO2H and alphaCys114-SOH. We replaced alphaGln90, which is conserved in all known NHases and involved in the hydrogen-bond network around the catalytic center, with glutamic acid or asparagine. The kcat of alphaQ90E and alphaQ90N mutants decreased to 24% and 5% that of wild type respectively, but the effect of mutations on Km was not very significant. In both mutants, the alphaCys114-SOH modification appeared to be responsible for the catalysis as in native NHase. We crystallized the nitrosylated alphaQ90N mutant and determined its structure at a resolution of 1.43 A. The structure was basically identical to that of native nitrosylated NHase except for the mutated site and its vicinity. The structural difference between native and alphaQ90N mutant NHases suggested the importance of the hydrogen bond networks between alphaGln90 and the iron center for the catalytic activity.
- United Graduate School of Agricultural Science, Tokyo University of Agriculture and Technology, Fuchu, Tokyo, Japan.
Organizational Affiliation: