Solution structure of the SHP-1 C-terminal SH2 domain complexed with a tyrosine-phosphorylated peptide from NKG2A
Kasai, T., Koshiba, S., Inoue, M., Kigawa, T., Yokoyama, S.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Tyrosine-protein phosphatase non-receptor type 6 | 118 | Homo sapiens | Mutation(s): 0 Gene Names: SHP1 EC: 3.1.3.48 | 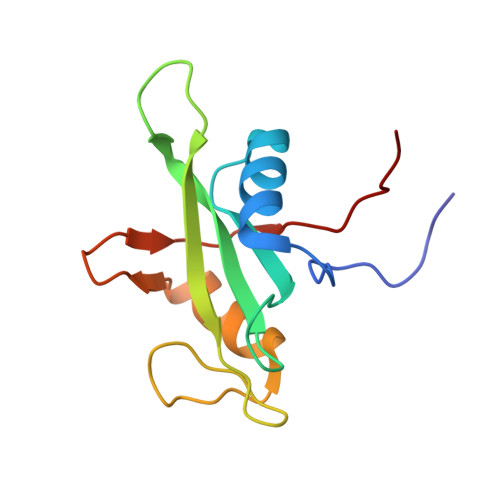 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P29350 GTEx: ENSG00000111679 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P29350 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| natural killer group 2A | 15 | N/A | Mutation(s): 0 | 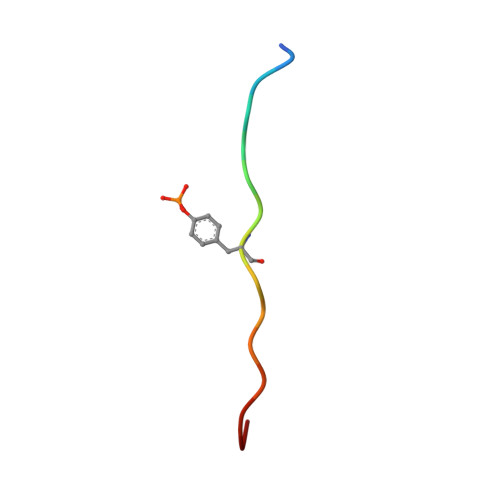 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P26715 GTEx: ENSG00000134545 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P26715 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| PTR Query on PTR | B | L-PEPTIDE LINKING | C9 H12 N O6 P |  | TYR |