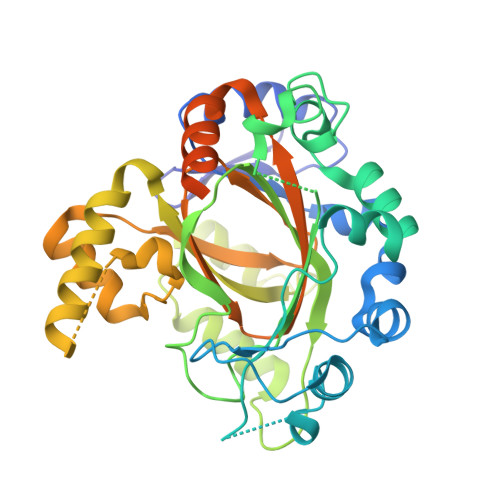
Crystal Structure of the Jumonji Domain of Human Jumonji Domain Containing 1C Protein
Vollmar, M., Johansson, C., Krojer, T., Strain-Damerell, C., Froese, S., Williams, E., Coutandin, D., von Delft, F., Weigelt, J., Arrowsmith, C.H., Bountra, C., Edwards, A., Oppermann, U.To be published.