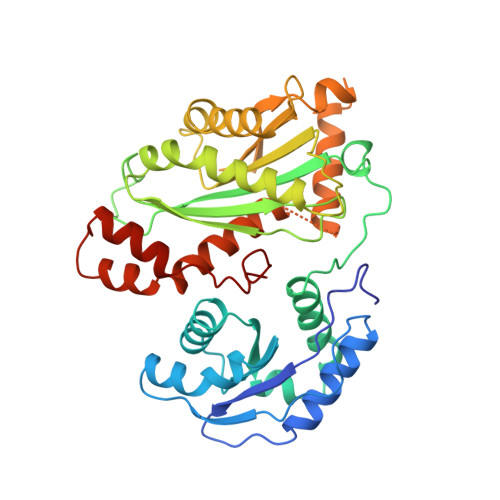
Crystal Structure of the Mid-Piwi Lobe of a Eukaryotic Argonaute Protein
Boland, A., Huntzinger, E., Schmidt, S., Izaurralde, E., Weichenriede, O.(2011) Proc Natl Acad Sci U S A 108: 10466
- PubMed: 21646546 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1103946108
- Primary Citation Related Structures:
2YHA, 2YHB - PubMed Abstract:
Argonaute proteins (AGOs) are essential effectors in RNA-mediated gene silencing pathways. They are characterized by a bilobal architecture, in which one lobe contains the N-terminal and PAZ domains and the other contains the MID and PIWI domains. Here, we present the first crystal structure of the MID-PIWI lobe from a eukaryotic AGO, the Neurospora crassa QDE-2 protein. Compared to prokaryotic AGOs, the domain orientation is conserved, indicating a conserved mode of nucleic acid binding. The PIWI domain shows an adaptable surface loop next to a eukaryote-specific α-helical insertion, which are both likely to contact the PAZ domain in a conformation-dependent manner to sense the functional state of the protein. The MID-PIWI interface is hydrophilic and buries residues that were previously thought to participate directly in the allosteric regulation of guide RNA binding. The interface includes the binding pocket for the guide RNA 5' end, and residues from both domains contribute to binding. Accordingly, micro-RNA (miRNA) binding is particularly sensitive to alteration in the MID-PIWI interface in Drosophila melanogaster AGO1 in vivo. The structure of the QDE-2 MID-PIWI lobe provides molecular and mechanistic insight into eukaryotic AGOs and has significant implications for understanding the role of these proteins in silencing.
- Max Planck Institute for Developmental Biology, Spemannstrasse 35, D-72076 Tübingen, Germany.
Organizational Affiliation: