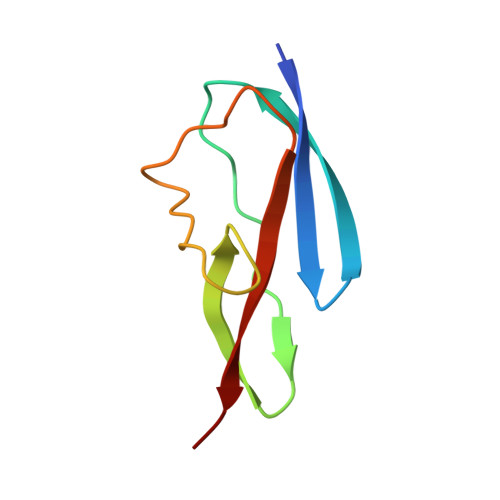
Fold and Function of the Inlb B-Repeat.
Ebbes, M., Bleymuller, W.M., Cernescu, M., Nolker, R., Brutschy, B., Niemann, H.H.(2011) J Biological Chem 286: 15496
- PubMed: 21345802 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.189951
- Primary Citation Related Structures:
2Y5P, 2Y5Q - PubMed Abstract:
Host cell invasion by the facultative intracellular pathogen Listeria monocytogenes requires the invasion protein InlB in many cell types. InlB consists of an N-terminal internalin domain that binds the host cell receptor tyrosine kinase Met and C-terminal GW domains that bind to glycosaminoglycans (GAGs). Met binding and activation is required for host cell invasion, while the interaction between GW domains and GAGs enhances this effect. Soluble InlB elicits the same cellular phenotypes as the natural Met ligand hepatocyte growth factor/scatter factor (HGF/SF), e.g. cell scatter. So far, little is known about the central part of InlB, the B-repeat. Here we present a structural and functional characterization of the InlB B-repeat. The crystal structure reveals a variation of the β-grasp fold that is most similar to small ubiquitin-like modifiers (SUMOs). However, structural similarity also suggests a potential evolutionary relation to bacterial mucin-binding proteins. The B-repeat defines the prototype structure of a hitherto uncharacterized domain present in over a thousand bacterial proteins. Generally, this domain probably acts as a spacer or a receptor-binding domain in extracellular multi-domain proteins. In cellular assays the B-repeat acts synergistically with the internalin domain conferring to it the ability to stimulate cell motility. Thus, the B-repeat probably binds a further host cell receptor and thereby enhances signaling downstream of Met.
- Department of Chemistry, Bielefeld University, Universitätsstrasse 25, 33615 Bielefeld, Germany.
Organizational Affiliation: