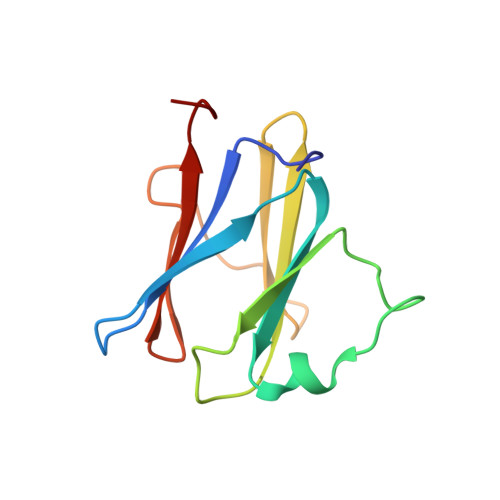
Structural and Functional Studies on the N-Terminal Domain of the Shigella Type III Secretion Protein Mxig.
Mcdowell, M.A., Johnson, S., Deane, J.E., Cheung, M., Roehrich, A.D., Blocker, A.J., Mcdonnell, J.M., Lea, S.M.(2011) J Biological Chem 286: 30606
- PubMed: 21733840 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.243865
- Primary Citation Related Structures:
2XXS - PubMed Abstract:
MxiG is a single-pass membrane protein that oligomerizes within the inner membrane ring of the Shigella flexneri type III secretion system (T3SS). The MxiG N-terminal domain (MxiG-N) is the predominant cytoplasmic structure; however, its role in T3SS assembly and secretion is largely uncharacterized. We have determined the solution structure of MxiG-N residues 6-112 (MxiG-N(6-112)), representing the first published structure of this T3SS domain. The structure shows strong structural homology to forkhead-associated (FHA) domains. Canonically, these cell-signaling modules bind phosphothreonine (Thr(P)) via highly conserved residues. However, the putative phosphate-binding pocket of MxiG-N(6-112) does not align with other FHA domain structures or interact with Thr(P). Furthermore, mutagenesis of potential phosphate-binding residues has no effect on S. flexneri T3SS assembly and function. Therefore, MxiG-N has a novel function for an FHA domain. Positioning of MxiG-N(6-112) within the EM density of the S. flexneri needle complex gives insight into the ambiguous stoichiometry of the T3SS, supporting models with 24 MxiG subunits in the inner membrane ring.
- Sir William Dunn School of Pathology, University of Oxford, Oxford OX1 3RE, United Kingdom.
Organizational Affiliation: