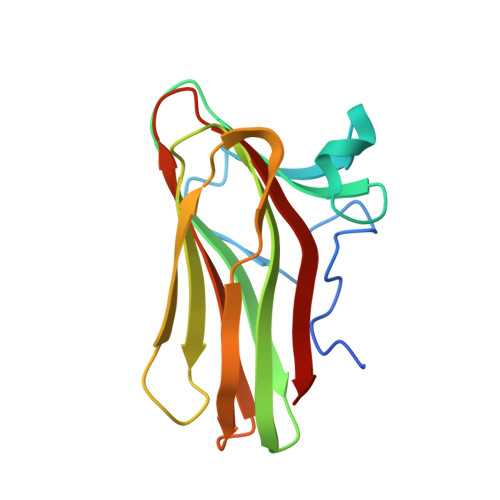
Recognition of the Helical Structure of Beta-1,4-Galactan by a New Family of Carbohydrate-Binding Modules.
Cid, M., Pedersen, H.L., Kaneko, S., Coutinho, P.M., Henrissat, B., Willats, W.G.T., Boraston, A.B.(2010) J Biological Chem 285: 35999
- PubMed: 20826814 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.166330
- Primary Citation Related Structures:
2XOM, 2XON - PubMed Abstract:
The microbial enzymes that depolymerize plant cell wall polysaccharides, ultimately promoting energy liberation and carbon recycling, are typically complex in their modularity and often contain carbohydrate-binding modules (CBMs). Here, through analysis of an unknown module from a Thermotoga maritima endo-β-1,4-galactanase, we identify a new family of CBMs that are most frequently found appended to proteins with β-1,4-galactanase activity. Polysaccharide microarray screening, immunofluorescence microscopy, and biochemical analysis of the isolated module demonstrate the specificity of the module, here called TmCBM61, for β-1,4-linked galactose-containing ligands, making it the founding member of family CBM61. The ultra-high resolution X-ray crystal structures of TmCBM61 (0.95 and 1.4 Å resolution) in complex with β-1,4-galactotriose reveal the molecular basis of the specificity of the CBM for β-1,4-galactan. Analysis of these structures provides insight into the recognition of an unexpected helical galactan conformation through a mode of binding that resembles the recognition of starch.
- Department Biochemistry and Microbiology, University of Victoria, Victoria, British Columbia V8W 3P6, Canada.
Organizational Affiliation: