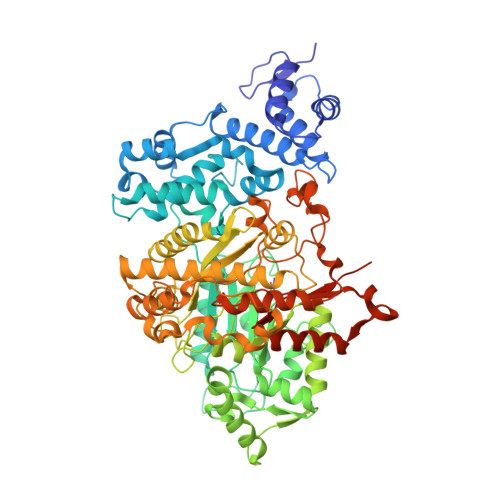
Kinetics of Radical Intermediate Formation and Deoxynucleotide Production in 3-Aminotyrosine- Substituted Escherichia Coli Ribonucleotide Reductases.
Minnihan, E.C., Seyedsayamdost, M.R., Uhlin, U., Stubbe, J.(2011) J Am Chem Soc 133: 9430
- PubMed: 21612216 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja201640n
- Primary Citation Related Structures:
2XO4, 2XO5 - PubMed Abstract:
Escherichia coli ribonucleotide reductase is an α2β2 complex and catalyzes the conversion of nucleoside 5'-diphosphates (NDPs) to 2'-deoxynucleotides (dNDPs). The reaction is initiated by the transient oxidation of an active-site cysteine (C(439)) in α2 by a stable diferric tyrosyl radical (Y(122)•) cofactor in β2. This oxidation occurs by a mechanism of long-range proton-coupled electron transfer (PCET) over 35 Å through a specific pathway of residues: Y(122)•→ W(48)→ Y(356) in β2 to Y(731)→ Y(730)→ C(439) in α2. To study the details of this process, 3-aminotyrosine (NH(2)Y) has been site-specifically incorporated in place of Y(356) of β. The resulting protein, Y(356)NH(2)Y-β2, and the previously generated proteins Y(731)NH(2)Y-α2 and Y(730)NH(2)Y-α2 (NH(2)Y-RNRs) are shown to catalyze dNDP production in the presence of the second subunit, substrate (S), and allosteric effector (E) with turnover numbers of 0.2-0.7 s(-1). Evidence acquired by three different methods indicates that the catalytic activity is inherent to NH(2)Y-RNRs and not the result of copurifying wt enzyme. The kinetics of formation of 3-aminotyrosyl radical (NH(2)Y•) at position 356, 731, and 730 have been measured with all S/E pairs. In all cases, NH(2)Y• formation is biphasic (k(fast) of 9-46 s(-1) and k(slow) of 1.5-5.0 s(-1)) and kinetically competent to be an intermediate in nucleotide reduction. The slow phase is proposed to report on the conformational gating of NH(2)Y• formation, while the k(cat) of ~0.5 s(-1) is proposed to be associated with rate-limiting oxidation by NH(2)Y• of the subsequent amino acid on the pathway during forward PCET. The X-ray crystal structures of Y(730)NH(2)Y-α2 and Y(731)NH(2)Y-α2 have been solved and indicate minimal structural changes relative to wt-α2. From the data, a kinetic model for PCET along the radical propagation pathway is proposed.
- Department of Chemistry, Massachusetts Institute of Technology, 77 Massachusetts Avenue, Cambridge, Massachusetts 02139, USA.
Organizational Affiliation: