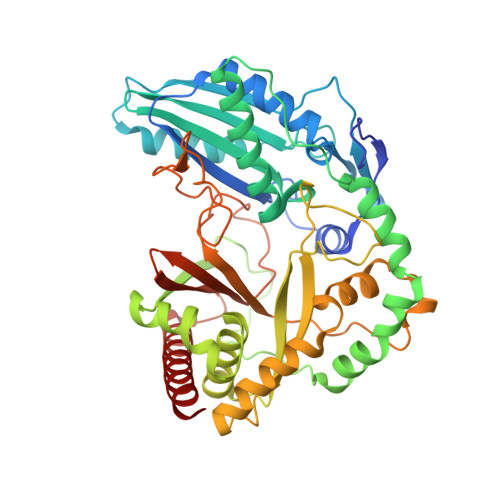
Structure of the Epimerization Domain of Tyrocidine Synthetase A
Samel, S.-A., Czodrowski, P., Essen, L.-O.(2014) Acta Crystallogr D Biol Crystallogr 70: 1442
- PubMed: 24816112 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004714004398
- Primary Citation Related Structures:
2XHG - PubMed Abstract:
Tyrocidine, a macrocyclic decapeptide from Bacillus brevis, is nonribosomally assembled by a set of multimodular peptide synthetases, which condense two D-amino acids and eight L-amino acids to produce this membrane-disturbing antibiotic. D-Phenylalanine, the first amino acid incorporated into tyrocidine, is catalytically derived from enzyme-bound L-Phe by the C-terminal epimerization (E) domain of tyrocidine synthetase A (TycA). The 1.5 Å resolution structure of the cofactor-independent TycA E domain reveals an intimate relationship to the condensation (C) domains of peptide synthetases. In contrast to the latter, the TycA E domain uses an enlarged bridge region to plug the active-site canyon from the acceptor side, whereas at the donor side a latch-like floor loop is suitably extended to accommodate the αIII helix of the preceding peptide-carrier domain. Additionally, E domains exclusively harbour a conserved glutamate residue, Glu882, that is opposite the active-site residue His743. This active-site topology implies Glu882 as a candidate acid-base catalyst, whereas His743 stabilizes in the protonated state a transient enolate intermediate of the L↔D isomerization.
- Biomedical Research Centre/FB15, Philipps Universität, Hans-Meerwein-Strasse 4, 35032 Marburg, Germany.
Organizational Affiliation: