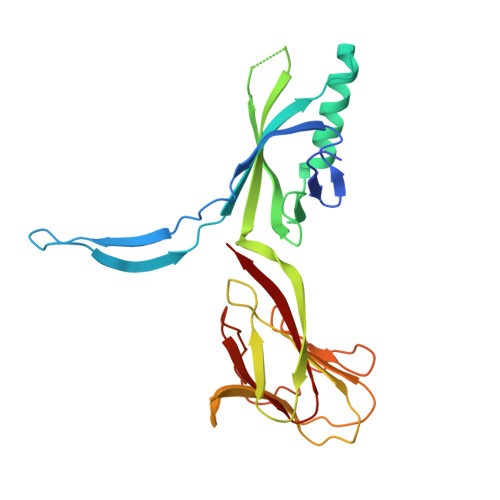
Crystal Structure of Bacteriophage Spp1 Distal Tail Protein (Gp 19.1): A Baseplate Hub Paradigm in Gram+ Infecting Phages.
Veesler, D., Robin, G., Lichiere, J., Auzat, I., Tavares, P., Bron, P., Campanacci, V., Cambillau, C.(2010) J Biological Chem 285: 36666
- PubMed: 20843802 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.157529
- Primary Citation Related Structures:
2X8K - PubMed Abstract:
Siphophage SPP1 infects the gram-positive bacterium Bacillus subtilis using its long non-contractile tail and tail-tip. Electron microscopy (EM) previously allowed a low resolution assignment of most orf products belonging to these regions. We report here the structure of the SPP1 distal tail protein (Dit, gp19.1). The combination of x-ray crystallography, EM, and light scattering established that Dit is a back-to-back dimer of hexamers. However, Dit fitting in the virion EM maps was only possible with a hexamer located between the tail-tube and the tail-tip. Structure comparison revealed high similarity between Dit and a central component of lactophage baseplates. Sequence similarity search expanded its relatedness to several phage proteins, suggesting that Dit is a docking platform for the tail adsorption apparatus in Siphoviridae infecting gram-positive bacteria and that its architecture is a paradigm for these hub proteins. Dit structural similarity extends also to non-contractile and contractile phage tail proteins (gpV(N) and XkdM) as well as to components of the bacterial type 6 secretion system, supporting an evolutionary connection between all these devices.
- Architecture et Fonction des Macromolécules Biologiques, UMR 6098 CNRS and Universités d'Aix-Marseille I & II, Campus de Luminy, Case 932, 13288 Marseille Cedex 09, France.
Organizational Affiliation: