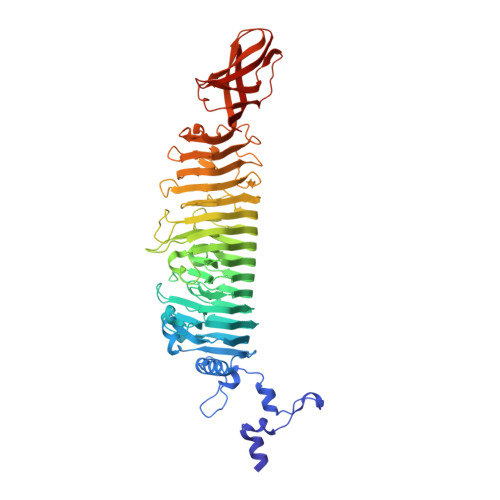
Single Amino Acid Exchange in Bacteriophage Hk620 Tailspike Protein Results in Thousand-Fold Increase of its Oligosaccharide Affinity.
Broeker, N.K., Gohlke, U., Muller, J.J., Uetrecht, C., Heinemann, U., Seckler, R., Barbirz, S.(2013) Glycobiology 23: 59
- PubMed: 22923442 Search on PubMed
- DOI: https://doi.org/10.1093/glycob/cws126
- Primary Citation Related Structures:
2X6W, 2X6X, 2X6Y, 2X85, 4AVZ - PubMed Abstract:
Bacteriophage HK620 recognizes and cleaves the O-antigen polysaccharide of Escherichia coli serogroup O18A1 with its tailspike protein (TSP). HK620TSP binds hexasaccharide fragments with low affinity, but single amino acid exchanges generated a set of high-affinity mutants with submicromolar dissociation constants. Isothermal titration calorimetry showed that only small amounts of heat were released upon complex formation via a large number of direct and solvent-mediated hydrogen bonds between carbohydrate and protein. At room temperature, association was both enthalpy- and entropy-driven emphasizing major solvent rearrangements upon complex formation. Crystal structure analysis showed identical protein and sugar conformers in the TSP complexes regardless of their hexasaccharide affinity. Only in one case, a TSP mutant bound a different hexasaccharide conformer. The extended sugar binding site could be dissected in two regions: first, a hydrophobic pocket at the reducing end with minor affinity contributions. Access to this site could be blocked by a single aspartate to asparagine exchange without major loss in hexasaccharide affinity. Second, a region where the specific exchange of glutamate for glutamine created a site for an additional water molecule. Side-chain rearrangements upon sugar binding led to desolvation and additional hydrogen bonding which define this region of the binding site as the high-affinity scaffold.
- Physikalische Biochemie, Universität Potsdam, Germany.
Organizational Affiliation: