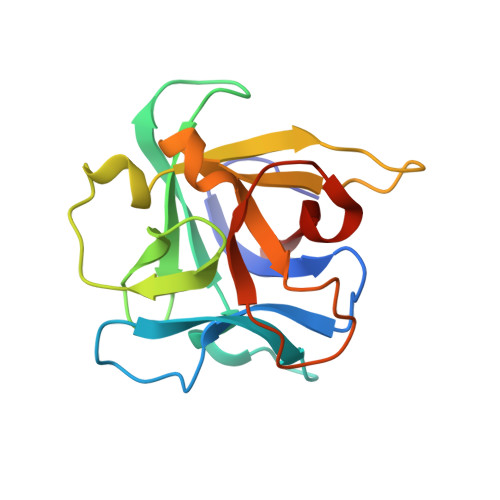
Crystal Structure of the Galnac/Gal-Specific Agglutinin from the Phytopathogenic Ascomycete Sclerotinia Sclerotiorum Reveals Novel Adaptation of a Beta-Trefoil Domain
Sulzenbacher, G., Roig-Zamboni, V., Peumans, W.J., Rouge, P., Van Damme, E.J.M., Bourne, Y.(2010) J Mol Biology 400: 715
- PubMed: 20566411 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2010.05.038
- Primary Citation Related Structures:
2X2S, 2X2T - PubMed Abstract:
A lectin from the phytopathogenic ascomycete Sclerotinia sclerotiorum that shares only weak sequence similarity with characterized fungal lectins has recently been identified. S. sclerotiorum agglutinin (SSA) is a homodimeric protein consisting of two identical subunits of approximately 17 kDa and displays specificity primarily towards Gal/GalNAc. Glycan array screening indicates that SSA readily interacts with Gal/GalNAc-bearing glycan chains. The crystal structures of SSA in the ligand-free form and in complex with the Gal-beta1,3-GalNAc (T-antigen) disaccharide have been determined at 1.6 and 1.97 A resolution, respectively. SSA adopts a beta-trefoil domain as previously identified for other carbohydrate-binding proteins of the ricin B-like lectin superfamily and accommodates terminal non-reducing galactosyl and N-acetylgalactosaminyl glycans. Unlike other structurally related lectins, SSA contains a single carbohydrate-binding site at site alpha. SSA reveals a novel dimeric assembly markedly dissimilar to those described earlier for ricin-type lectins. The present structure exemplifies the adaptability of the beta-trefoil domain in the evolution of fungal lectins.
- Architecture et Fonction des Macromolécules Biologiques UMR-6098 CNRS, Université Aix-Marseille, Campus Luminy, Case 932, F-13288 Marseille cedex 09, France.
Organizational Affiliation: