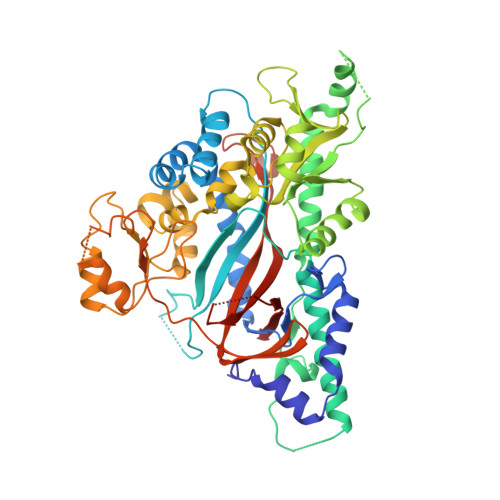
Structure of Trypanosoma Brucei Glutathione Synthetase; Domain and Loop Alterations in the Catalytic Cycle of a Highly Conserved Enzyme.
Fyfe, P.K., Alphey, M.S., Hunter, W.N.(2010) Mol Biochem Parasitol 170: 93
- PubMed: 20045436 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molbiopara.2009.12.011
- Primary Citation Related Structures:
2WYO - PubMed Abstract:
Glutathione synthetase catalyses the synthesis of the low molecular mass thiol glutathione from l-gamma-glutamyl-l-cysteine and glycine. We report the crystal structure of the dimeric enzyme from Trypanosoma brucei in complex with the product glutathione. The enzyme belongs to the ATP-grasp family, a group of enzymes known to undergo conformational changes upon ligand binding. The T. brucei enzyme crystal structure presents two dimers in the asymmetric unit. The structure reveals variability in the order and position of a small domain, which forms a lid for the active site and serves to capture conformations likely to exist during the catalytic cycle. Comparisons with orthologous enzymes, in particular from Homo sapiens and Saccharomyces cerevisae, indicate a high degree of sequence and structure conservation in part of the active site. Structural differences that are observed between the orthologous enzymes are assigned to different ligand binding states since key residues are conserved. This suggests that the molecular determinants of ligand recognition and reactivity are highly conserved across species. We conclude that it would be difficult to target the parasite enzyme in preference to the host enzyme and therefore glutathione synthetase may not be a suitable target for antiparasitic drug discovery.
- Division of Biological Chemistry and Drug Discovery, College of Life Sciences, University of Dundee, Dow Street, Dundee, DD1 5EH, United Kingdom.
Organizational Affiliation: