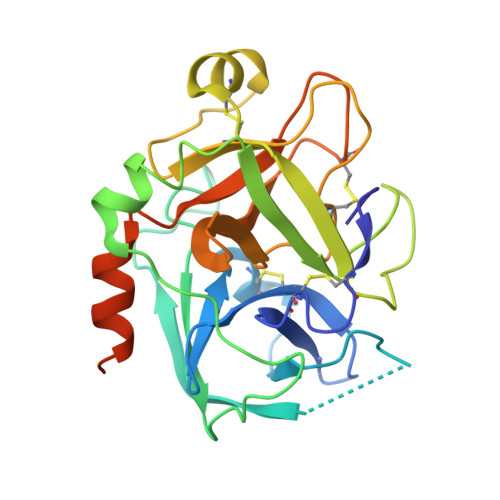
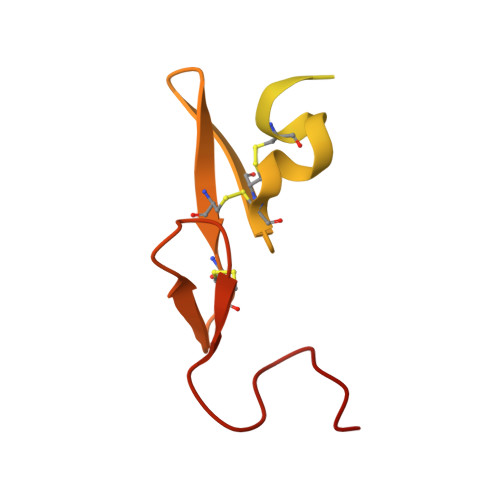
Structure and Property Based Design of Factor Xa Inhibitors: Pyrrolidin-2-Ones with Monoaryl P4 Motifs
Kleanthous, S., Borthwick, A.D., Brown, D., Burns-Kurtis, C.L., Campbell, M., Chaudry, L., Chan, C., Clarte, M., Convery, M.A., Harling, J.D., Hortense, E., Irving, W.R., Irvine, S., Pateman, A.J., Patikis, A., Pinto, I.L., Pollard, D.R., Roethka, T.J., Senger, S., Shah, G.P., Stelman, G.J., Toomey, J.R., Watson, N.S., Whittaker, C., Zhou, P., Young, R.J.(2010) Bioorg Med Chem Lett 20: 618
- PubMed: 20006499 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2009.11.077
- Primary Citation Related Structures:
2WYG, 2WYJ - PubMed Abstract:
Structure and property based drug design was exploited in the synthesis of sulfonamidopyrrolidin-2-one-based factor Xa inhibitors, incorporating neutral and basic monoaryl P4 groups, ultimately producing potent inhibitors with effective levels of anticoagulant activity and extended oral pharmacokinetic profiles. However, time dependant inhibition of Cytochrome P450 3A4 was a particular issue with this series.
- GlaxoSmithKline, Medicines Research Centre, Gunnels Wood Road, Stevenage, Hertfordshire SG1 2NY, United Kingdom.
Organizational Affiliation: