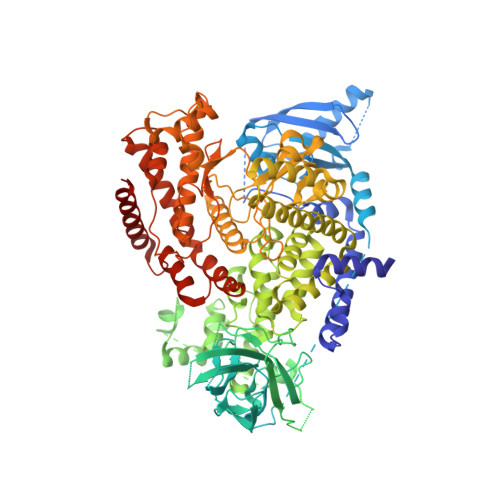
The P110D Structure: Mechanisms for Selectivity and Potency of New Pi(3)K Inhibitors
Berndt, A., Miller, S., Williams, O., Lee, D.D., Houseman, B.T., Pacold, J.I., Gorrec, F., Hon, W.-C., Liu, Y., Rommel, C., Gaillard, P., Ruckle, T., Schwarz, M.K., Shokat, K.M., Shaw, J.P., Williams, R.L.(2010) Nat Chem Biol 6: 117
- PubMed: 20081827 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.293
- Primary Citation Related Structures:
2WXF, 2WXG, 2WXH, 2WXI, 2WXJ, 2WXK, 2WXL, 2WXM, 2WXN, 2WXO, 2WXP, 2WXQ, 2WXR, 2X38 - PubMed Abstract:
Deregulation of the phosphoinositide-3-OH kinase (PI(3)K) pathway has been implicated in numerous pathologies including cancer, diabetes, thrombosis, rheumatoid arthritis and asthma. Recently, small-molecule and ATP-competitive PI(3)K inhibitors with a wide range of selectivities have entered clinical development. In order to understand the mechanisms underlying the isoform selectivity of these inhibitors, we developed a new expression strategy that enabled us to determine to our knowledge the first crystal structure of the catalytic subunit of the class IA PI(3)K p110 delta. Structures of this enzyme in complex with a broad panel of isoform- and pan-selective class I PI(3)K inhibitors reveal that selectivity toward p110 delta can be achieved by exploiting its conformational flexibility and the sequence diversity of active site residues that do not contact ATP. We have used these observations to rationalize and synthesize highly selective inhibitors for p110 delta with greatly improved potencies.
- Medical Research Council-Laboratory of Molecular Biology, Cambridge, UK.
Organizational Affiliation: