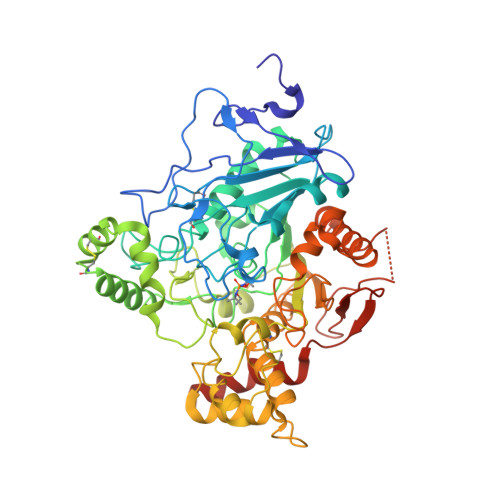
Crystal Structures of Oxime-Bound Fenamiphos-Acetylcholinesterases: Reactivation Involving Flipping of the His447 Ring to Form a Reactive Glu334-His447-Oxime Triad.
Hornberg, A., Artursson, E., Warme, R., Pang, Y.-P., Ekstrom, F.(2010) Biochem Pharmacol 79: 507
- PubMed: 19732756 Search on PubMed
- DOI: https://doi.org/10.1016/j.bcp.2009.08.027
- Primary Citation Related Structures:
2WU3, 2WU4 - PubMed Abstract:
Organophosphorus insecticides and nerve agents inhibit the vital enzyme acetylcholinesterase by covalently bonding to the catalytic serine residue of the enzyme. Oxime-based reactivators, such as [(E)-[1-[(4-carbamoylpyridin-1-ium-1-yl)methoxymethyl]pyridin-2-ylidene]methyl]-oxoazanium dichloride (HI-6) and 1,7-heptylene-bis-N,N'-2-pyridiniumaldoxime dichloride (Ortho-7), restore the organophosphate-inhibited enzymatic activity by cleaving the phosphorous conjugate. In this article, we report the intermolecular interactions between Mus musculus acetylcholinesterase inhibited by the insecticide fenamiphos (fep-mAChE) and HI-6 or Ortho-7 revealed by a combination of crystallography and kinetics. The crystal structures of the two oxime-bound fep-mAChE complexes show that both oximes interact with the peripheral anionic site involving different conformations of Trp286 and different peripheral-site residues (Tyr124 for HI-6 and Tyr72 for Ortho-7). Moreover, residues at catalytic site of the HI-6-bound fep-mAChE complex adopt conformations that are similar to those in the apo mAChE, whereas significant conformational changes are observed for the corresponding residues in the Ortho-7-bound fep-mAChE complex. Interestingly, flipping of the His447 imidazole ring allows the formation of a hydrogen bonding network among the Glu334-His447-Ortho-7 triad, which presumably deprotonates the Ortho-7 oxime hydroxyl group, increases the nucleophilicity of the oxime group, and leads to cleavage of the phosphorous conjugate. These results offer insights into a detailed reactivation mechanism for the oximes and development of improved reactivators.
- Swedish Defence Research Agency, Division of CBRN Defence and Security, S-901 82 Umeå, Sweden.
Organizational Affiliation: