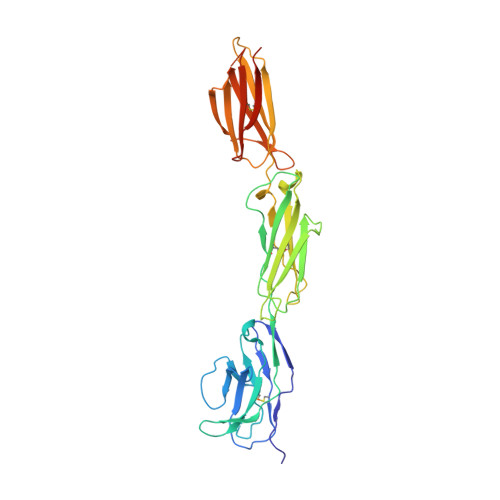
Structure of Signal-Regulatory Protein Alpha: A Link to Antigen Receptor Evolution.
Hatherley, D., Graham, S.C., Harlos, K., Stuart, D.I., Barclay, A.N.(2009) J Biological Chem 284: 26613
- PubMed: 19628875 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.017566
- Primary Citation Related Structures:
2WNG - PubMed Abstract:
Signal-regulatory protein alpha (SIRPalpha) is a myeloid membrane receptor that interacts with the membrane protein CD47, a marker of self. We have solved the structure of the complete extracellular portion of SIRPalpha, comprising three immunoglobulin superfamily domains, by x-ray crystallography to 2.5 A resolution. These data, together with previous data on the N-terminal domain and its ligand CD47 (possessing a single immunoglobulin superfamily domain), show that the CD47-SIRPalpha interaction will span a distance of around 14 nm between interacting cells, comparable with that of an immunological synapse. The N-terminal (V-set) domain mediates binding to CD47, and the two others are found to be constant (C1-set) domains. C1-set domains are restricted to proteins involved in vertebrate antigen recognition: T cell antigen receptors, immunoglobulins, major histocompatibility complex antigens, tapasin, and beta2-microglobulin. The domains of SIRPalpha (domains 2 and 3) are structurally more similar to C1-set domains than any cell surface protein not involved in antigen recognition. This strengthens the suggestion from sequence analysis that SIRP is evolutionarily closely related to antigen recognition proteins.
- Sir William Dunn School of Pathology, University of Oxford, Oxford OX1 3RE, United Kingdom.
Organizational Affiliation: