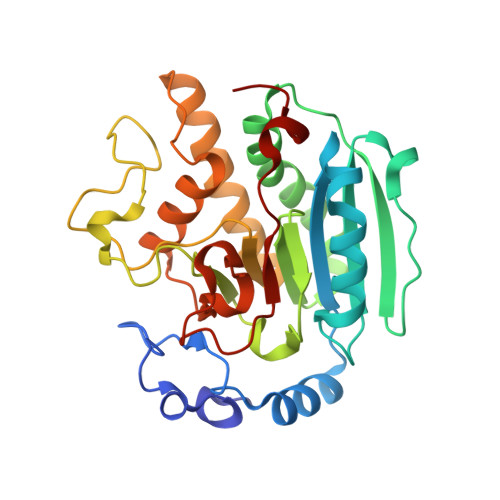
Crystal Structure of Alpha-1,3 Galactosyltransferase (Alpha3Gt) in a Complex with P-Nitrophenyl-Beta-Galactoside (Pnpbetagal).
Jamaluddin, H., Tumbale, P., Ferns, T.A., Thiyagarajan, N., Brew, K., Acharya, K.R.(2009) Biochem Biophys Res Commun 385: 601
- PubMed: 19486884 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2009.05.111
- Primary Citation Related Structures:
2WGZ - PubMed Abstract:
The specificities of glycosyltransferases make them useful for the synthesis of biologically active oligosaccharides, but also restrict their range of products. In substrate engineering, substrate promiscuity is enhanced by attaching removable interactive groups to weak substrates. Thus, the attachment of betap-nitrophenyl converts galactose from a poor into a good substrate of alpha-1,3-galactosyltransferase. The crystallographic structure of a complex of alpha3GT containing p-nitrophenyl-beta-galactoside shows that the p-nitrophenyl binds similarly to the N-acetylglucosamine of the substrate, N-acetyllactosamine, interacting with the indole of Trp249. p-Nitrophenyl, unlike N-acetylglucosamine, makes no H-bonds but has more non-polar interactions, making it an effective monosaccharide mimetic.
- Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA27AY, UK.
Organizational Affiliation: