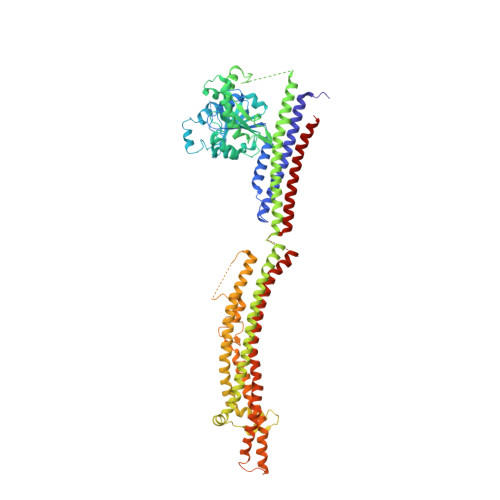
Structure of a Bacterial Dynamin-Like Protein Lipid Tube Provides a Mechanism for Assembly and Membrane Curving.
Low, H.H., Sachse, C., Amos, L.A., Lowe, J.(2009) Cell 139: 1342-1352
- PubMed: 20064379 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2009.11.003
- Primary Citation Related Structures:
2W6D - PubMed Abstract:
Proteins of the dynamin superfamily mediate membrane fission, fusion, and restructuring events by polymerizing upon lipid bilayers and forcing regions of high curvature. In this work, we show the electron cryomicroscopy reconstruction of a bacterial dynamin-like protein (BDLP) helical filament decorating a lipid tube at approximately 11 A resolution. We fitted the BDLP crystal structure and produced a molecular model for the entire filament. The BDLP GTPase domain dimerizes and forms the tube surface, the GTPase effector domain (GED) mediates self-assembly, and the paddle region contacts the lipids and promotes curvature. Association of BDLP with GMPPNP and lipid induces radical, large-scale conformational changes affecting polymerization. Nucleotide hydrolysis seems therefore to be coupled to polymer disassembly and dissociation from lipid, rather than membrane restructuring. Observed structural similarities with rat dynamin 1 suggest that our results have broad implication for other dynamin family members.
- MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 0QH, UK. h.low@mail.cryst.bbk.ac.uk
Organizational Affiliation: