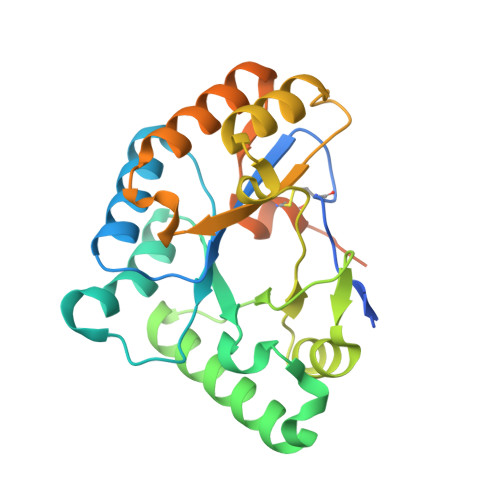
Structural and Functional Characterization of a Putative Polysaccharide Deacetylase of the Human Parasite Encephalitozoon Cuniculi.
Urch, J.E., Hurtado-Guerrero, R., Brosson, D., Liu, Z., Eijsink, V.G.H., Texier, C., Van Aalten, D.M.F.(2009) Protein Sci 18: 1197
- PubMed: 19472335 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.128
- Primary Citation Related Structures:
2VYO - PubMed Abstract:
The microsporidian Encephalitozoon cuniculi is an intracellular eukaryotic parasite considered to be an emerging opportunistic human pathogen. The infectious stage of this parasite is a unicellular spore that is surrounded by a chitin containing endospore layer and an external proteinaceous exospore. A putative chitin deacetylase (ECU11_0510) localizes to the interface between the plasma membrane and the endospore. Chitin deacetylases are family 4 carbohydrate esterases in the CAZY classification, and several bacterial members of this family are involved in evading lysis by host glycosidases, through partial de-N-acetylation of cell wall peptidoglycan. Similarly, ECU11_0510 could be important for E. cuniculi survival in the host, by protecting the chitin layer from hydrolysis by human chitinases. Here, we describe the biochemical, structural, and glycan binding properties of the protein. Enzymatic analyses showed that the putative deacetylase is unable to deacetylate chitooligosaccharides or crystalline beta-chitin. Furthermore, carbohydrate microarray analysis revealed that the protein bound neither chitooligosaccharides nor any of a wide range of other glycans or chitin. The high resolution crystal structure revealed dramatic rearrangements in the positions of catalytic and substrate binding residues, which explain the loss of deacetylase activity, adding to the unusual structural plasticity observed in other members of this esterase family. Thus, it appears that the ECU11_0510 protein is not a carbohydrate deacetylase and may fulfill an as yet undiscovered role in the E. cuniculi parasite.
- Division of Molecular Microbiology, College of Life Sciences, University of Dundee, Dundee, Scotland.
Organizational Affiliation: