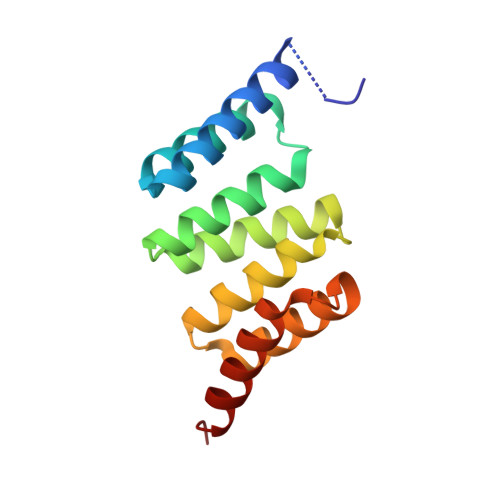
Structural and Functional Characterization of Human Sgt and its Interaction with Vpu of the Human Immunodeficiency Virus Type 1.
Dutta, S., Tan, Y.J.(2008) Biochemistry 47: 10123
- PubMed: 18759457 Search on PubMed
- DOI: https://doi.org/10.1021/bi800758a
- Primary Citation Related Structures:
2VYI - PubMed Abstract:
The small glutamine-rich tetratricopeptide repeat protein (SGT) belongs to a family of cochaperones that interacts with both Hsp70 and Hsp90 via the so-called TPR domain. Here, we present the crystal structure of the TPR domain of human SGT (SGT-TPR), which shows that it contains typical features found in the structures of other TPR domains. Previous studies show that full-length SGT can bind to both Vpu and Gag of human immunodeficiency virus type 1 (HIV-1) and the overexpression of SGT in cells reduces the efficiency of HIV-1 particle release. We show that SGT-TPR can bind Vpu and reduce the amount of HIV-1 p24, which is the viral capsid, secreted from cells transfected with the HIV-1 proviral construct, albeit at a lower efficiency than full-length SGT. This indicates that the TPR domain of SGT is sufficient for the inhibition of HIV-1 particle release but the N- and/or C-terminus also have some contributions. The SGT binding site in Vpu was also identified by using peptide array and confirmed by GST pull-down assay.
- Collaborative Antiviral Research Group, Cancer and Developmental Cell Biology Division, Institute of Molecular and Cell Biology, Agency for Science, Technology and Research, 61 Biopolis Drive, Singapore 138673.
Organizational Affiliation: