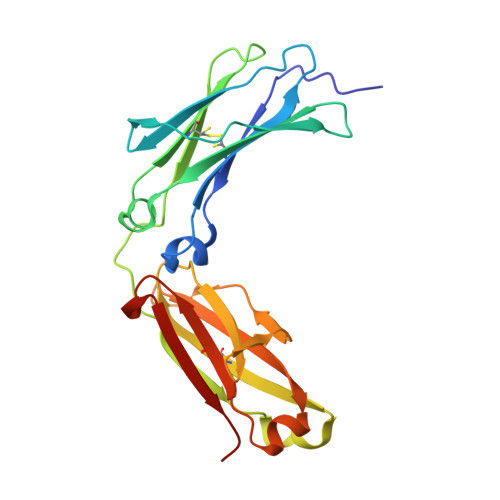
The Crystal Structure of Rabbit Igg-Fc.
Girardi, E., Holdom, M.D., Davies, A.M., Sutton, B.J., Beavil, A.J.(2009) Biochem J 417: 77
- PubMed: 18764781 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20081355
- Primary Citation Related Structures:
2VUO - PubMed Abstract:
We report the structure of the Fc fragment of rabbit IgG at 1.95 A (1 A=0.1 nm) resolution. Rabbit IgG was the molecule for which Porter established the four-chain, Upsilon-shaped structure of the antibody molecule, and crystals of the Fc ('Fragment crystallisable') were first reported almost 50 years ago in this journal [Porter, R. R. (1959) Biochem. J. 73, 119-126]. This high-resolution analysis, apparently of the same crystal form, reveals several features of IgG-Fc structure that have not previously been described. More of the lower hinge region is visible in this structure than in others, demonstrating not only the acute bend in the IgG molecule that this region can mediate, as seen in receptor complexes, but also that this region has a tendency to adopt a bent structure even in the absence of receptor. As observed in other IgG-Fc structures, the Cgamma2 domains display greater mobility/disorder within the crystals than the Cgamma3 domains; unexpectedly the structure reveals partial cleavage of both Cgamma2 intra-domain disulphide bonds, whereas an alternative conformation for one of the cysteine residues in the intact bridge within the more ordered Cgamma3 domains is observed. The N-linked oligosaccharide chains at Asn(297) are well-defined and reveal two alternative conformations for the galactose units on each of the alpha(1-6)-linked branches. The presence of this galactose unit is important for stabilizing the structure of the entire branched carbohydrate chain, and its absence correlates with the severity of autoimmune conditions such as rheumatoid arthritis in both human clinical studies and in a rabbit model of the disease. Rabbit IgG, through this high-resolution structure of its Fc region, thus continues to offer new insights into antibody structure.
- Randall Division of Cell and Molecular Biophysics, King's College London, New Hunt's House, Guy's Campus, London SE11UL, UK.
Organizational Affiliation: