Potent Memapsin 2 (Beta-Secretase) Inhibitors: Design, Synthesis, Protein-Ligand X-Ray Structure, and in Vivo Evaluation.
Ghosh, A.K., Kumaragurubaran, N., Hong, L., Kulkarni, S., Xu, X., Miller, H.B., Reddy, D.S., Weerasena, V., Turner, R., Chang, W., Koelsch, G., Tang, J.(2008) Bioorg Med Chem Lett 18: 1031
- PubMed: 18180160 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bmcl.2007.12.028
- Primary Citation Related Structures:
2VKM - PubMed Abstract:
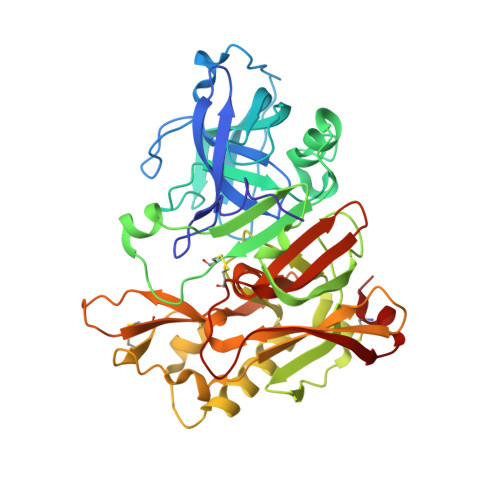
Structure-based design, synthesis, and biological evaluation of a series of peptidomimetic beta-secretase inhibitors incorporating hydroxyethylamine isosteres are described. We have identified inhibitor 24 which has shown exceedingly potent activity in memapsin 2 enzyme inhibitory (K(i) 1.8 nM) and cellular (IC(50)=1 nM in Chinese hamster ovary cells) assays. Inhibitor 24 has also shown very impressive in vivo properties (up to 65% reduction of plasma A beta) in transgenic mice. The X-ray structure of protein-ligand complex of memapsin 2 revealed critical interactions in the memapsin 2 active site.
- Department of Chemistry, Purdue University, West Lafayette, IN 47907, USA. akghosh@purdue.edu
Organizational Affiliation: