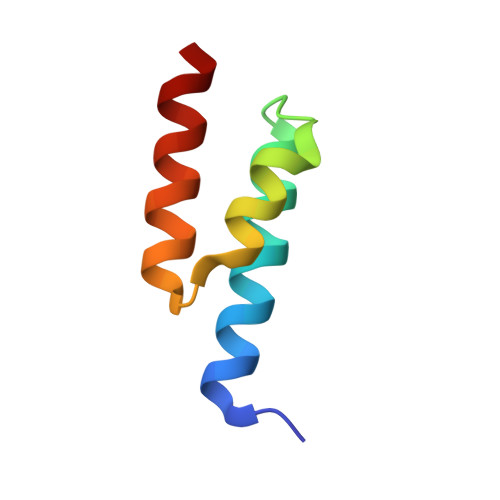
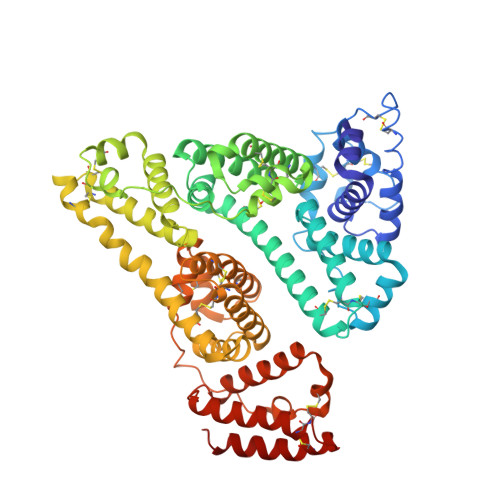
Structural Basis for the Binding of Naproxen to Human Serum Albumin in the Presence of Fatty Acids and the Ga Module.
Lejon, S., Cramer, J.F., Nordberg, P.A.(2008) Acta Crystallogr Sect F Struct Biol Cryst Commun 64: 64
- PubMed: 18259051 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S174430910706770X
- Primary Citation Related Structures:
2VDB - PubMed Abstract:
The previously determined crystal structure of the bacterial albumin-binding GA module in complex with human serum albumin (HSA) suggested the possibility of utilizing the complex in the study of ligand binding to HSA. As a continuation of these studies, the crystal structure of the HSA-GA complex with the drug molecule naproxen and the fatty acid decanoate bound to HSA has been determined to a resolution of 2.5 A. In terms of drug binding, the structure suggests that the binding of decanoate to the albumin molecule may play a role in making the haemin site in subdomain IB of the albumin molecule available for the binding of naproxen. In addition, structure comparisons with solved structures of HSA and of the HSA-GA complex show that the GA module is capable of binding to different conformations of HSA. The HSA-GA complex therefore emerges as a possible platform for the crystallographic study of specific HSA-drug interactions and of the influence exerted by the presence of fatty acids.
- Department of Cell and Molecular Biology, Uppsala University, Biomedical Centre, Box 596, S-751 24 Uppsala, Sweden. sara@xray.bmc.uu.se
Organizational Affiliation: