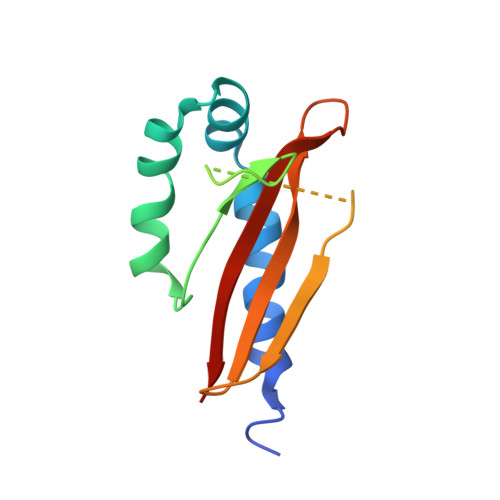
A Ligand-Induced Switch in the Periplasmic Domain of Sensor Histidine Kinase Cita.
Sevvana, M., Vijayan, V., Zweckstetter, M., Reinelt, S., Madden, D.R., Herbst-Irmer, R., Sheldrick, G.M., Bott, M., Griesinger, C., Becker, S.(2008) J Mol Biology 377: 512
- PubMed: 18258261 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.01.024
- Primary Citation Related Structures:
2J80, 2V9A - PubMed Abstract:
Sensor histidine kinases of two-component signal-transduction systems are essential for bacteria to adapt to variable environmental conditions. However, despite their prevalence, it is not well understood how extracellular signals such as ligand binding regulate the activity of these sensor kinases. CitA is the sensor histidine kinase in Klebsiella pneumoniae that regulates the transport and anaerobic metabolism of citrate in response to its extracellular concentration. We report here the X-ray structures of the periplasmic sensor domain of CitA in the citrate-free and citrate-bound states. A comparison of the two structures shows that ligand binding causes a considerable contraction of the sensor domain. This contraction may represent the molecular switch that activates transmembrane signaling in the receptor.
- Department of Structural Chemistry, University of Göttingen, Tammannstrasse 4, 37077 Göttingen, Germany.
Organizational Affiliation: