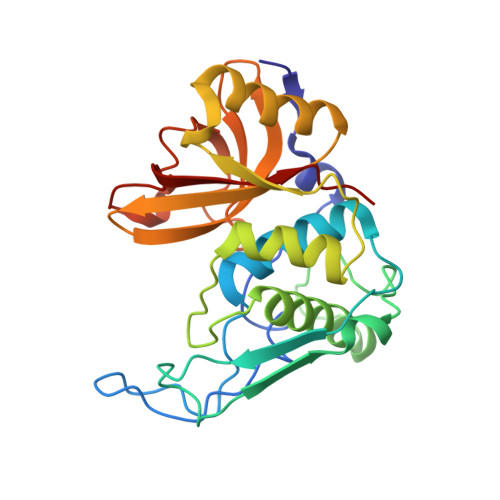
Structure of the Mature Streptococcal Cysteine Protease Exotoxin Mspeb in its Active Dimeric Form.
Olsen, J.G., Dagil, R., Niclasen, L.M., Soerensen, O.E., Kragelund, B.B.(2009) J Mol Biology 393: 693
- PubMed: 19712682 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.08.046
- Primary Citation Related Structures:
2UZJ - PubMed Abstract:
Invasive infections of Streptococcus pyogenes are dependent on the cysteine protease streptococcal pyrogenic exotoxin B. Previous structures of the enzyme have not disclosed the proper active-site configuration. Here, the crystal structure of the mature enzyme is presented to 1.55 A, disclosing a homodimer. A serine from one subunit inserts into the active site of the other to donate to the oxyanion hole and coordinates the ligand proximal to the active-site cysteine. Dimerization is unique to the mature form and is clearly a prerequisite for catalysis. The present structure supports a tripartite switch system that is triggered upon dimerization and substrate binding: (1) liberation of the active-site histidine from an inactive configuration, (2) relocation of residues blocking the substrate binding pockets and (3) repositioning of two active-site tryptophans to settle in the active configuration. Based on the present structure, the active site of clan CA cysteine proteases is expanded and a detailed mechanism of the deacylation mechanism is proposed. The results may have applications for the development of protease inhibitors specific to bacterial cysteine proteases.
- Structural Biology and NMR Laboratory, Department of Biology, University of Copenhagen, Ole Maaloes Vej 5, Copenhagen, Denmark.
Organizational Affiliation: