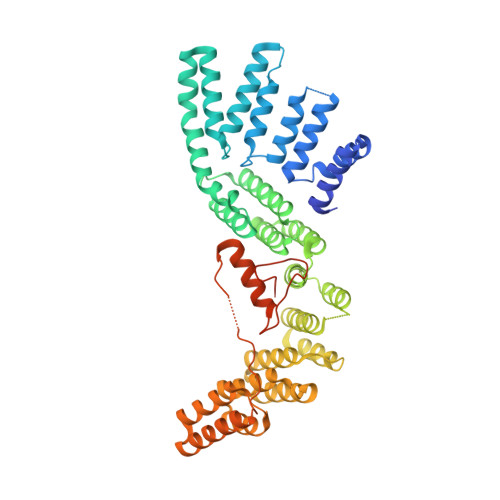
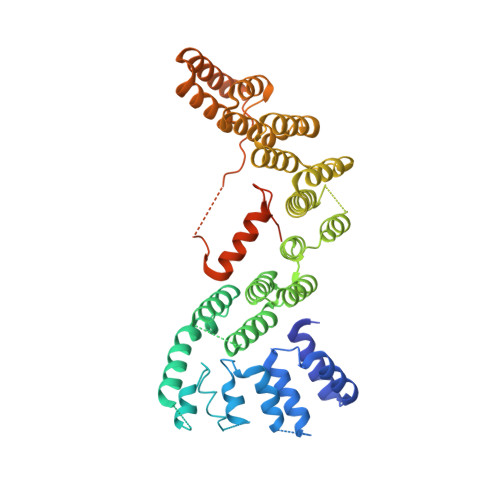
The Structure of the Cstf-77 Homodimer Provides Insights Into Cstf Assembly.
Legrand, P., Pinaud, N., Minvielle-Sebastia, L., Fribourg, S.(2007) Nucleic Acids Res 35: 4515
- PubMed: 17584787 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkm458
- Primary Citation Related Structures:
2UY1 - PubMed Abstract:
The cleavage stimulation factor (CstF) is essential for the first step of poly(A) tail formation at the 3' ends of mRNAs. This heterotrimeric complex is built around the 77-kDa protein bridging both CstF-64 and CstF-50 subunits. We have solved the crystal structure of the 77-kDa protein from Encephalitozoon cuniculi at a resolution of 2 A. The structure folds around 11 Half-a-TPR repeats defining two domains. The crystal structure reveals a tight homodimer exposing phylogenetically conserved areas for interaction with protein partners. Mapping experiments identify the C-terminal region of Rna14p, the yeast counterpart of CstF-77, as the docking domain for Rna15p, the yeast CstF-64 homologue.
- Institut Européen de Chimie et Biologie, INSERM U869, 2 rue Robert Escarpit Pessac, F-33607, Université Victor Segalen, Bordeaux 2.
Organizational Affiliation: