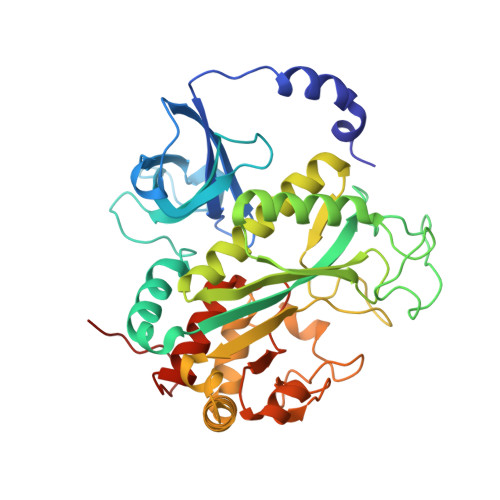
Crystal Structures of Mammalian Glutamine Synthetases Illustrate Substrate-Induced Conformational Changes and Provide Opportunities for Drug and Herbicide Design.
Krajewski, W.W., Collins, R., Holmberg-Schiavone, L., Jones, T.A., Karlberg, T., Mowbray, S.L.(2008) J Mol Biology 375: 217
- PubMed: 18005987 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.10.029
- Primary Citation Related Structures:
2OJW, 2QC8, 2UU7 - PubMed Abstract:
Glutamine synthetase (GS) catalyzes the ligation of glutamate and ammonia to form glutamine, with concomitant hydrolysis of ATP. In mammals, the activity eliminates cytotoxic ammonia, at the same time converting neurotoxic glutamate to harmless glutamine; there are a number of links between changes in GS activity and neurodegenerative disorders, such as Alzheimer's disease. In plants, because of its importance in the assimilation and re-assimilation of ammonia, the enzyme is a target of some herbicides. GS is also a central component of bacterial nitrogen metabolism and a potential drug target. Previous studies had investigated the structures of bacterial and plant GSs. In the present publication, we report the first structures of mammalian GSs. The apo form of the canine enzyme was solved by molecular replacement and refined at a resolution of 3 A. Two structures of human glutamine synthetase represent complexes with: a) phosphate, ADP, and manganese, and b) a phosphorylated form of the inhibitor methionine sulfoximine, ADP and manganese; these structures were refined to resolutions of 2.05 A and 2.6 A, respectively. Loop movements near the active site generate more closed forms of the eukaryotic enzymes when substrates are bound; the largest changes are associated with the binding of the nucleotide. Comparisons with earlier structures provide a basis for the design of drugs that are specifically directed at either human or bacterial enzymes. The site of binding the amino acid substrate is highly conserved in bacterial and eukaryotic GSs, whereas the nucleotide binding site varies to a much larger degree. Thus, the latter site offers the best target for specific drug design. Differences between mammalian and plant enzymes are much more subtle, suggesting that herbicides targeting GS must be designed with caution.
- Department of Cell and Molecular Biology, Biomedical Center, Uppsala University, Box 596, SE-751 24 Uppsala, Sweden.
Organizational Affiliation: