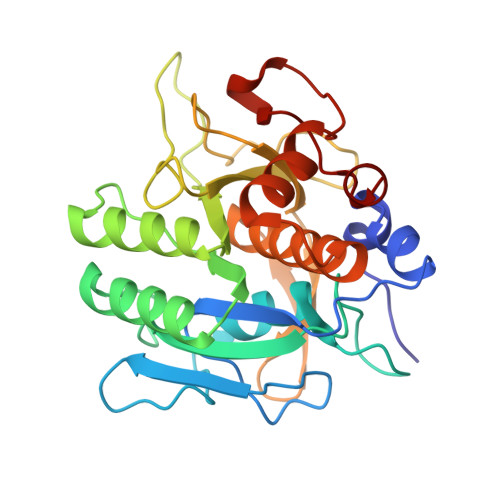
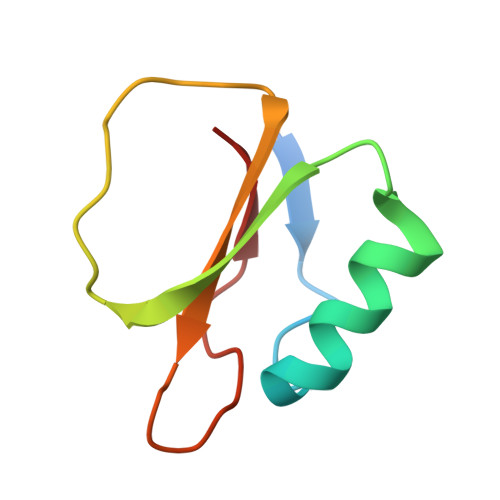
Structural comparison of two serine proteinase-protein inhibitor complexes: eglin-c-subtilisin Carlsberg and CI-2-subtilisin Novo.
McPhalen, C.A., James, M.N.(1988) Biochemistry 27: 6582-6598
- PubMed: 3064813 Search on PubMed
- Primary Citation Related Structures:
2SEC, 2SNI - PubMed Abstract:
The crystal structures of the molecular complexes between two serine proteinases and two of their protein inhibitors have been determined: subtilisin Carlsberg with the recombinant form of eglin-c from the leech Hirudo medicinalis and subtilisin Novo with chymotrypsin inhibitor 2 from barley seeds. The structures have been fully refined by restrained-parameter least-squares methods to crystallographic R factors (sigma[[Fo[ - [Fc[[/sigma[Fo[) of 0.136 at 1.8-A resolution and 0.154 at 2.1-A resolution, respectively. The 274 equivalent alpha-carbon atoms of the enzymes superpose with an rms deviation of 0.53 A. Sequence changes between the enzymes result in localized structural adjustments. Functional groups in the active sites superpose with an rms deviation of 0.19 A for 161 equivalent atoms; this close similarity in the conformation of active-site residues provides no obvious reason for known differences in catalytic activity between Carlsberg and Novo. Conformational changes in the active-site region indicate a small induced fit of enzyme and inhibitor. Some conformational differences are observed between equivalent active-site residues of subtilisin Carlsberg and alpha-chymotrypsin. Despite differences in tertiary architecture, most enzyme-substrate (inhibitor) interactions are maintained. Subtilisin Carlsberg has a rare cis-peptide bond preceding Thr211 (Gly211 in Novo). Both enzymes contain tightly bound Ca2+ ions. Site 1 is heptacoordinate with the oxygen atoms at the vertices of a pentagonal bipyramid. Site 2 in Carlsberg is probably occupied by a K+ ion in Novo. Conserved water molecules appear to play important structural roles in the enzyme interior, in the inhibitor beta-sheet, and at the enzyme-inhibitor interface. The 62 equivalent alpha-carbon atoms of the inhibitors superpose with an rms deviation of 1.68 A. Sequence changes result in somewhat different packing of the alpha-helix, beta-sheet, and reactive-site loop relative to each other. Hydrogen bonds and electrostatic interactions supporting the conformation of the reactive-site loop are conserved. The 24 main-chain plus C beta atoms of P4 to P1' overlap with an rms deviation of 0.19 A. Features contributing to the inhibitory nature of eglin-c and CI-2 are discussed.
- Department of Biochemistry, University of Alberta, Edmonton, Canada.
Organizational Affiliation: