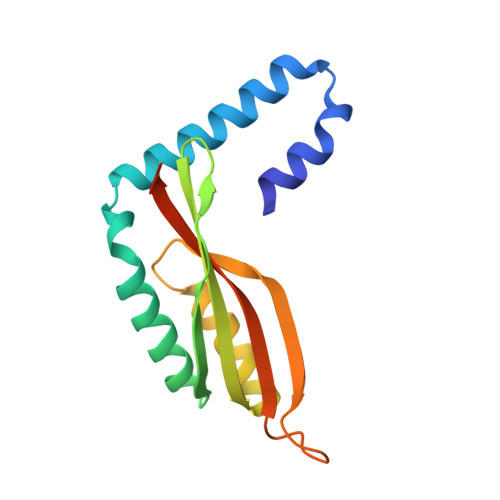
Periplasmic domains of Pseudomonas aeruginosa PilN and PilO form a stable heterodimeric complex.
Sampaleanu, L.M., Bonanno, J.B., Ayers, M., Koo, J., Tammam, S., Burley, S.K., Almo, S.C., Burrows, L.L., Howell, P.L.(2009) J Mol Biology 394: 143-159
- PubMed: 19857646 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.09.037
- Primary Citation Related Structures:
2RJZ - PubMed Abstract:
Type IV pili (T4P) are bacterial virulence factors responsible for attachment to surfaces and for twitching motility, a motion that involves a succession of pilus extension and retraction cycles. In the opportunistic pathogen Pseudomonas aeruginosa, the PilM/N/O/P proteins are essential for T4P biogenesis, and genetic and biochemical analyses strongly suggest that they form an inner-membrane complex. Here, we show through co-expression and biochemical analysis that the periplasmic domains of PilN and PilO interact to form a heterodimer. The structure of residues 69-201 of the periplasmic domain of PilO was determined to 2.2 A resolution and reveals the presence of a homodimer in the asymmetric unit. Each monomer consists of two N-terminal coiled coils and a C-terminal ferredoxin-like domain. This structure was used to generate homology models of PilN and the PilN/O heterodimer. Our structural analysis suggests that in vivo PilN/O heterodimerization would require changes in the orientation of the first N-terminal coiled coil, which leads to two alternative models for the role of the transmembrane domains in the PilN/O interaction. Analysis of PilN/O orthologues in the type II secretion system EpsL/M revealed significant similarities in their secondary structures and the tertiary structures of PilO and EpsM, although the way these proteins interact to form inner-membrane complexes appears to be different in T4P and type II secretion. Our analysis suggests that PilN interacts directly, via its N-terminal tail, with the cytoplasmic protein PilM. This work shows a direct interaction between the periplasmic domains of PilN and PilO, with PilO playing a key role in the proper folding of PilN. Our results suggest that PilN/O heterodimers form the foundation of the inner-membrane PilM/N/O/P complex, which is critical for the assembly of a functional T4P complex.
- Program in Molecular Structure and Function, Hospital for Sick Children, 555 University Avenue, Toronto, Ontario, Canada M5G 1X8.
Organizational Affiliation: