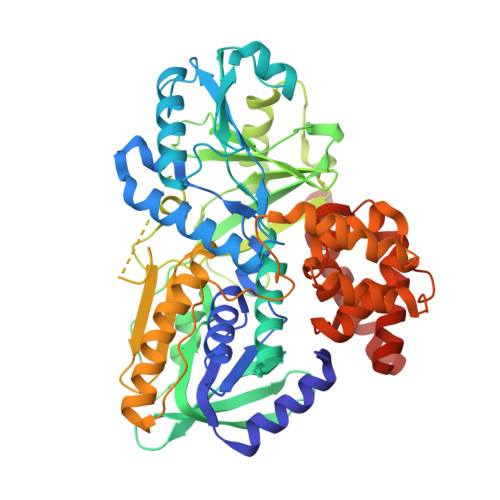
Structure of alpha-glycerophosphate oxidase from Streptococcus sp.: a template for the mitochondrial alpha-glycerophosphate dehydrogenase.
Colussi, T., Parsonage, D., Boles, W., Matsuoka, T., Mallett, T.C., Karplus, P.A., Claiborne, A.(2008) Biochemistry 47: 965-977
- PubMed: 18154320 Search on PubMed
- DOI: https://doi.org/10.1021/bi701685u
- Primary Citation Related Structures:
2RGH, 2RGO - PubMed Abstract:
The FAD-dependent alpha-glycerophosphate oxidase (GlpO) from Enterococcus casseliflavus and Streptococcus sp. was originally studied as a soluble flavoprotein oxidase; surprisingly, the GlpO sequence is 30-43% identical to those of the alpha-glycerophosphate dehydrogenases (GlpDs) from mitochondrial and bacterial sources. The structure of a deletion mutant of Streptococcus sp. GlpO (GlpODelta, lacking a 50-residue insert that includes a flexible surface region) has been determined using multiwavelength anomalous dispersion data and refined at 2.3 A resolution. Using the GlpODelta structure as a search model, we have also determined the intact GlpO structure, as refined at 2.4 A resolution. The first two domains of the GlpO fold are most closely related to those of the flavoprotein glycine oxidase, where they function in FAD binding and substrate binding, respectively; the GlpO C-terminal domain consists of two helix bundles and is not closely related to any known structure. The flexible surface region in intact GlpO corresponds to a segment of missing electron density that links the substrate-binding domain to a betabetaalpha element of the FAD-binding domain. In accordance with earlier biochemical studies (stabilizations of the covalent FAD-N5-sulfite adduct and p-quinonoid form of 8-mercapto-FAD), Ile430-N, Thr431-N, and Thr431-OG are hydrogen bonded to FAD-O2alpha in GlpODelta, stabilizing the negative charge in these two modified flavins and facilitating transfer of a hydride to FAD-N5 (from Glp) as well. Active-site overlays with the glycine oxidase-N-acetylglycine and d-amino acid oxidase-d-alanine complexes demonstrate that Arg346 of GlpODelta is structurally equivalent to Arg302 and Arg285, respectively; in both cases, these residues interact directly with the amino acid substrate or inhibitor carboxylate. The structural and functional divergence between GlpO and the bacterial and mitochondrial GlpDs is also discussed.
- Center for Structural Biology, Wake Forest University School of Medicine, Winston-Salem, North Carolina 27157, USA.
Organizational Affiliation: